Dimensionality Reduction - Kernel PCA#
Notebook by:
Royi Avital RoyiAvital@fixelalgorithms.com
Revision History#
Version |
Date |
User |
Content / Changes |
|---|---|---|---|
1.0.000 |
13/04/2024 |
Royi Avital |
First version |
# Import Packages
# General Tools
import numpy as np
import scipy as sp
import pandas as pd
# Machine Learning
from sklearn.datasets import make_circles
from sklearn.decomposition import KernelPCA, PCA
# Miscellaneous
import math
import os
from platform import python_version
import random
import timeit
# Typing
from typing import Callable, Dict, List, Optional, Self, Set, Tuple, Union
# Visualization
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
# Jupyter
from IPython import get_ipython
from IPython.display import Image
from IPython.display import display
from ipywidgets import Dropdown, FloatSlider, interact, IntSlider, Layout, SelectionSlider
from ipywidgets import interact
Notations#
(?) Question to answer interactively.
(!) Simple task to add code for the notebook.
(@) Optional / Extra self practice.
(#) Note / Useful resource / Food for thought.
Code Notations:
someVar = 2; #<! Notation for a variable
vVector = np.random.rand(4) #<! Notation for 1D array
mMatrix = np.random.rand(4, 3) #<! Notation for 2D array
tTensor = np.random.rand(4, 3, 2, 3) #<! Notation for nD array (Tensor)
tuTuple = (1, 2, 3) #<! Notation for a tuple
lList = [1, 2, 3] #<! Notation for a list
dDict = {1: 3, 2: 2, 3: 1} #<! Notation for a dictionary
oObj = MyClass() #<! Notation for an object
dfData = pd.DataFrame() #<! Notation for a data frame
dsData = pd.Series() #<! Notation for a series
hObj = plt.Axes() #<! Notation for an object / handler / function handler
Code Exercise#
Single line fill
vallToFill = ???
Multi Line to Fill (At least one)
# You need to start writing
????
Section to Fill
#===========================Fill This===========================#
# 1. Explanation about what to do.
# !! Remarks to follow / take under consideration.
mX = ???
???
#===============================================================#
# Configuration
# %matplotlib inline
seedNum = 512
np.random.seed(seedNum)
random.seed(seedNum)
# Matplotlib default color palette
lMatPltLibclr = ['#1f77b4', '#ff7f0e', '#2ca02c', '#d62728', '#9467bd', '#8c564b', '#e377c2', '#7f7f7f', '#bcbd22', '#17becf']
# sns.set_theme() #>! Apply SeaBorn theme
runInGoogleColab = 'google.colab' in str(get_ipython())
# Constants
FIG_SIZE_DEF = (8, 8)
ELM_SIZE_DEF = 50
CLASS_COLOR = ('b', 'r')
EDGE_COLOR = 'k'
MARKER_SIZE_DEF = 10
LINE_WIDTH_DEF = 2
# Courses Packages
import sys
sys.path.append('../')
sys.path.append('../../')
sys.path.append('../../../')
from utils.DataVisualization import PlotScatterData
# General Auxiliary Functions
Dimensionality Reduction by Kernel - PCA#
The Kernel PCA is a specific case of applying non linear transformation on the features and then applying the PCA transform.
Utilizing the Kernel Trick in the PCA framework allows an efficient computation of the transforms which can be defined by a kernel.
This notebook demonstrates a simple use case of the Kernel PCA.
# Parameters
# Data
numCircles0 = 350
numCircles1 = 350
noiseLevel = 0.01
# Model
kernelType = 'rbf'
γ = 10.0
α = 0.1
numComp = 2
Generate / Load Data#
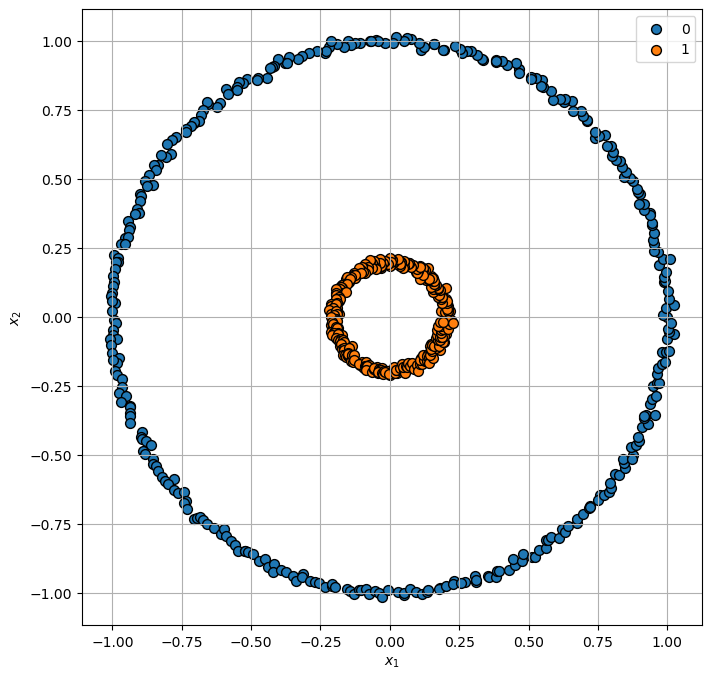
Generating concentric circles.
# Generate Data
numCircles = numCircles0 + numCircles1
mX, vY = make_circles((numCircles0, numCircles1), factor = 0.2, shuffle = False, noise = noiseLevel, random_state = seedNum)
numSamples = np.size(mX, 0)
print(f'The features data shape: {mX.shape}')
print(f'The labels data shape: {vY.shape}')
The features data shape: (700, 2)
The labels data shape: (700,)
Plot Data#
# Plot the Data
hF, hA = plt.subplots(figsize = (8, 8))
hA = PlotScatterData(mX, vY, hA)
Applying Dimensionality Reduction - Kernel PCA#
The Kernel-PCA is usually a framework used by other facilitators (MDS / IsoMap).
This section demonstrate the use of the Polynomial Kernel on the simple dataset.
# Applying the PCA Model
oPCA = PCA(n_components = numComp)
oPCA = oPCA.fit(mX)
# Applying the PCA Model
oKPCA = KernelPCA(n_components = numComp, kernel = kernelType, gamma = γ, fit_inverse_transform = True, alpha = α)
oKPCA = oKPCA.fit(mX)
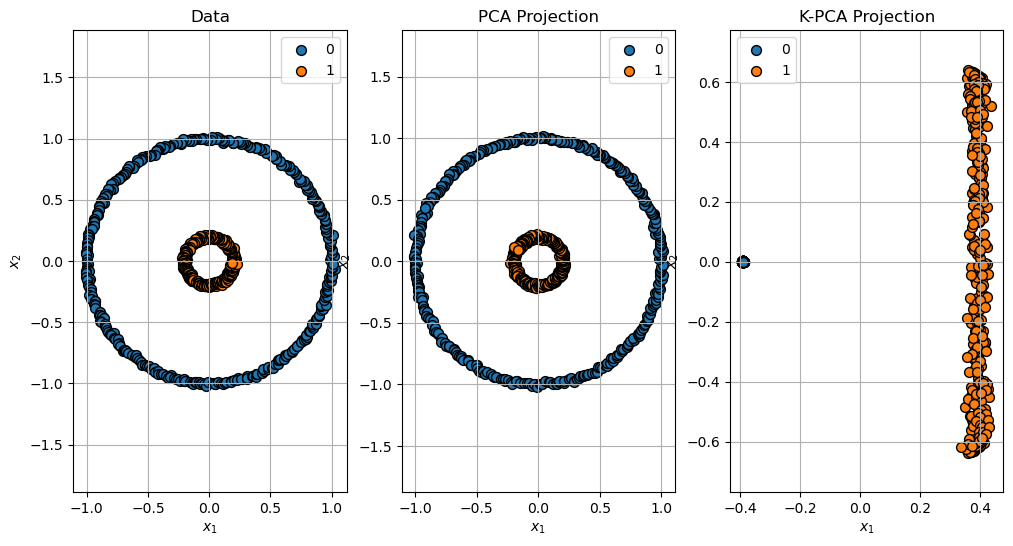
Projection of the Data#
Projection of the data: \(\mathbb{R}^{2} \to \mathbb{R}^{2}\).
# Projection
mXPca = oPCA.transform(mX)
mXKPca = oKPCA.transform(mX)
# Plot the Projection
hF, hA = plt.subplots(nrows = 1, ncols = 3, figsize = (12, 6))
hA[0] = PlotScatterData(mX, vY, hA[0])
hA[0].set_title('Data')
hA[0].axis('equal')
hA[1] = PlotScatterData(mXPca, vY, hA[1])
hA[1].set_title('PCA Projection')
hA[1].axis('equal')
hA[2] = PlotScatterData(mXKPca, vY, hA[2])
hA[2].set_title('K-PCA Projection')
hA[2].axis('equal')
plt.show()
Reconstruction of the Data#
# Reconstruction
mXPcaRec = oPCA.inverse_transform(mXPca)
mXKPcaRec = oKPCA.inverse_transform(mXKPca)
# Plot the Reconstruction
hF, hA = plt.subplots(nrows = 1, ncols = 3, figsize = (12, 6))
hA[0] = PlotScatterData(mX, vY, hA[0])
hA[0].set_title('Data')
hA[0].axis('equal')
hA[1] = PlotScatterData(mXPcaRec, vY, hA[1])
hA[1].set_title('PCA Reconstruction')
hA[1].axis('equal')
hA[2] = PlotScatterData(mXKPcaRec, vY, hA[2])
hA[2].set_title('K-PCA Reconstruction')
hA[2].axis('equal')
plt.show()
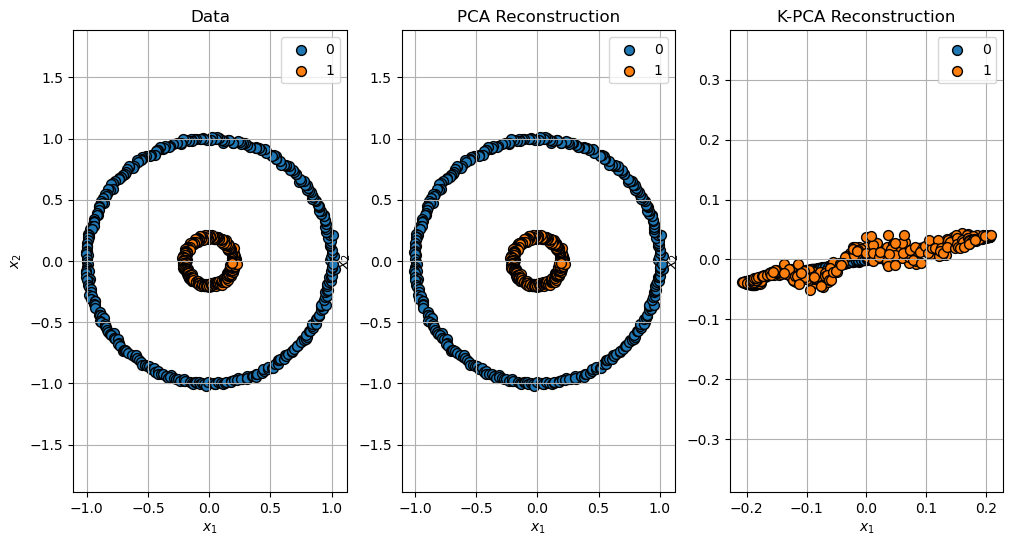
(?) Explain the reconstruction error of the K-PCA. You may read about the
fit_inverse_transformparameter.
