K-Means super pixcel#
Notebook by:
Royi Avital RoyiAvital@fixelalgorithms.com
Revision History#
Version |
Date |
User |
Content / Changes |
|---|---|---|---|
1.0.000 |
12/04/2024 |
Royi Avital |
First version |
# Import Packages
# General Tools
import numpy as np
import scipy as sp
import pandas as pd
# Image Processing Computer Vision
import skimage as ski
# Machine Learning
from kneed import KneeLocator #<! Elbow Method
from sklearn.base import BaseEstimator, ClusterMixin
from sklearn.preprocessing import MinMaxScaler
# Miscellaneous
import math
import os
from platform import python_version
import random
import timeit
# Typing
from typing import Callable, Dict, List, Optional, Self, Set, Tuple, Union
# Visualization
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
# Jupyter
from IPython import get_ipython
from IPython.display import Image
from IPython.display import display
from ipywidgets import Dropdown, FloatSlider, interact, IntSlider, Layout, SelectionSlider
from ipywidgets import interact
Notations#
(?) Question to answer interactively.
(!) Simple task to add code for the notebook.
(@) Optional / Extra self practice.
(#) Note / Useful resource / Food for thought.
Code Notations:
someVar = 2; #<! Notation for a variable
vVector = np.random.rand(4) #<! Notation for 1D array
mMatrix = np.random.rand(4, 3) #<! Notation for 2D array
tTensor = np.random.rand(4, 3, 2, 3) #<! Notation for nD array (Tensor)
tuTuple = (1, 2, 3) #<! Notation for a tuple
lList = [1, 2, 3] #<! Notation for a list
dDict = {1: 3, 2: 2, 3: 1} #<! Notation for a dictionary
oObj = MyClass() #<! Notation for an object
dfData = pd.DataFrame() #<! Notation for a data frame
dsData = pd.Series() #<! Notation for a series
hObj = plt.Axes() #<! Notation for an object / handler / function handler
Code Exercise#
Single line fill
vallToFill = ???
Multi Line to Fill (At least one)
# You need to start writing
????
Section to Fill
#===========================Fill This===========================#
# 1. Explanation about what to do.
# !! Remarks to follow / take under consideration.
mX = ???
???
#===============================================================#
# Configuration
# %matplotlib inline
seedNum = 512
np.random.seed(seedNum)
random.seed(seedNum)
# Matplotlib default color palette
lMatPltLibclr = ['#1f77b4', '#ff7f0e', '#2ca02c', '#d62728', '#9467bd', '#8c564b', '#e377c2', '#7f7f7f', '#bcbd22', '#17becf']
# sns.set_theme() #>! Apply SeaBorn theme
runInGoogleColab = 'google.colab' in str(get_ipython())
# Constants
FIG_SIZE_DEF = (8, 8)
ELM_SIZE_DEF = 50
CLASS_COLOR = ('b', 'r')
EDGE_COLOR = 'k'
MARKER_SIZE_DEF = 10
LINE_WIDTH_DEF = 2
# Courses Packages
# General Auxiliary Functions
def ConvertRgbToLab( mRgb: np.ndarray ) -> np.ndarray:
# Converts sets of RGB features into LAB features.
# Input (numPx x 3)
# Output: (numPx x 3)
mRgb3D = np.reshape(mRgb, (1, -1, 3))
mLab3D = ski.color.rgb2lab(mRgb3D)
return np.reshape(mLab3D, (-1, 3))
def PlotSuperPixels( mI: np.ndarray, mMask: np.ndarray, boundColor: Tuple[float, float, float] = (0.0, 1.0, 1.0), figSize: Tuple[int, int] = FIG_SIZE_DEF, hA: Optional[plt.Axes] = None ) -> plt.Axes:
if hA is None:
hF, hA = plt.subplots(figsize = figSize)
else:
hF = hA.get_figure()
mO = ski.segmentation.mark_boundaries(mI, mMask, boundColor)
hA.imshow(mO)
return hA
def PlotFeaturesHist( mX: np.ndarray, figSize: Tuple[int, int] = FIG_SIZE_DEF, hF: Optional[plt.Figure] = None ) -> plt.Figure:
numFeatures = mX.shape[1]
if hF is None:
hF, hA = plt.subplots(nrows = 1, ncols = numFeatures, figsize = figSize)
else:
hA = np.array(hF.axes)
hA = hA.flat
if len(hA) != numFeatures:
raise ValueError(f'The number of axes in the figure: {len(hA)} does not match the number of features: {numFeatures} in the data')
for ii in range(numFeatures):
hA[ii].hist(mX[:, ii])
return hF
Clustering by K-Means#
In this notebook we’ll do the following things:
Implement K-Means manually.
Use the K-Means to extract Super Pixels (A model for image segmentation).
The Super Pixel is a simple clustering method which says s Super Pixel is a cluster f pixels which are localized and have similar colors.
Hence it fits to be applied using a clustering method with the features being the values of the pixels and its coordinates.
The steps are as following:
Load the
Fruits.jpegimage and covert it to NumPy arraymIusing the SciKit Image library.
Seeimread().The image is given by \( I \in \mathbb{R}^{m \times n \times c}\) where \( c = \) is the number of channels (
RGB).
We need to convert it into \( X \in \mathbb{R}^{mn \times 3}\).
Namely, a 2D array where each row is the RGB values triplets.Feature Engineering:
Convert data into a color space with approximately euclidean metric -> LAB Color Space.
Add the Row / Column indices of each pixel as one the features.
Scale features to have the same range.
Apply K-Means clustering on the features.
Use the label of each pixel (The cluster it belongs to) to segment the image (Create Super Pixels).
Plot the segmentation (Super Pixels) map.
(#) You may try different color spaces.
(#) You may try different scaling of the features.

# Parameters
# Data
imgUrl = r'https://github.com/FixelAlgorithmsTeam/FixelCourses/raw/master/MachineLearningMethod/16_ParametricClustering/Fruits.jpeg'
# Model
numClusters = 50
numIter = 500
Generate / Load Data#
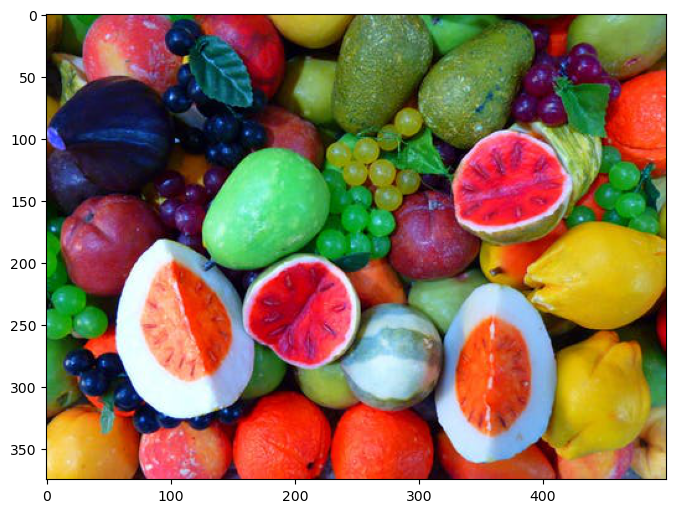
Load the fruits image.
# Load Data
mI = ski.io.imread(imgUrl)
print(f'The image shape: {mI.shape}')
The image shape: (375, 500, 3)
Plot Data#
# Plot the Data
hF, hA = plt.subplots(figsize = (8, 8))
# Display the image
hA.imshow(mI)
plt.show()

Pre Processing#
We need to convert the image from (numRows, numCols, numChannels) to (numRows * numCols, numChannels).
In our case, numChannels = 3 as we work with RGB image.
# Convert Image into Features Matrix
numRows, numCols, numChannel = mI.shape
#===========================Fill This===========================#
mX = np.reshape(mI, (numRows * numCols, numChannel))
#===============================================================#
print(f'The features data shape: {mX.shape}')
The features data shape: (187500, 3)
Feature Engineering#
In this section we’ll apply the feature engineering:
Convert data into a meaningful color space (LAB).
Add the location (Row / Column indices) information as a feature to segment the image.
Scale features to have similar dynamic range.
# Convert Features into LAB Color Space
mX = ConvertRgbToLab(mX) #<! Have a look on the function
# Add the Indices as Features
#===========================Fill This===========================#
# 1. Create a vector of the row index of each pixel.
# 2. Create a vector of the column index of each pixel.
# 3. Stack them as additional columns to `mX`.
# !! Python is column major.
# !! The number of elements in the vectors should match the number of pixels.
# !! You may find `repeat()` and `tile()` functions (NumPy) useful.
vR = np.repeat(np.arange(numRows), repeats = numCols) #<! Row indices
vC = np.tile(np.arange(numCols), reps = numRows) #<! Column indices
mX = np.column_stack((mX, vR, vC))
#===============================================================#
numFeat = mX.shape[1]
print(f'The features data shape: {mX.shape}')
The features data shape: (187500, 5)
# Plot the Features Histogram
hF, hA = plt.subplots(nrows = 1, ncols = numFeat, figsize = (15, 8))
hF = PlotFeaturesHist(mX, hF = hF)
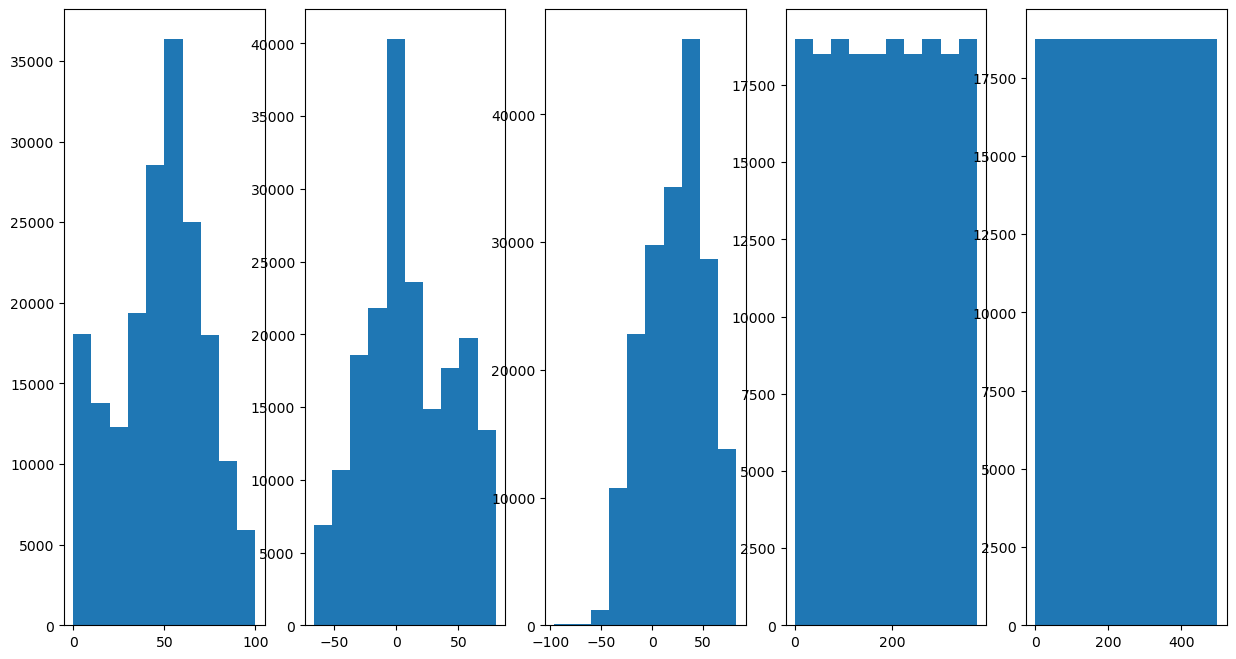
# Scale Features
# Scale each feature into the [0, 1] range.
# Having similar scaling means have similar contribution when calculating the distance.
#===========================Fill This===========================#
# 1. Construct the `MinMaxScaler` object.
# 2. Use it to transform the data.
oMinMaxScaler = MinMaxScaler()
mX = oMinMaxScaler.fit_transform(mX)
#===============================================================#
(?) What would happen if we didn’t scale the row and columns indices features?
# Plot the Features Histogram
hF, hA = plt.subplots(nrows = 1, ncols = numFeat, figsize = (15, 8))
hF = PlotFeaturesHist(mX, hF = hF)
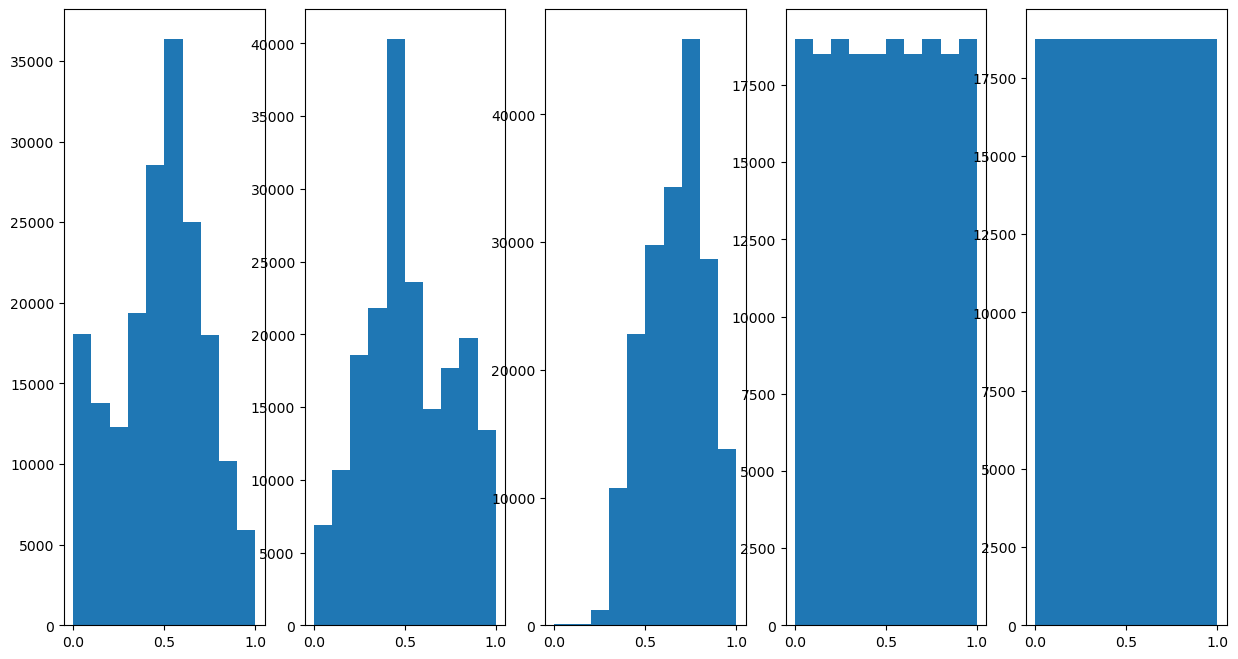
Cluster Data by K-Means#
Step I:
Assume fixed centroids \(\left\{ \boldsymbol{\mu}_{k}\right\} \), find the optimal clusters \(\left\{ \mathcal{D}_{k}\right\} \):
$\(\arg\min_{\left\{ \mathcal{D}_{k}\right\} }\sum_{k = 1}^{K}\sum_{\boldsymbol{x}_{i}\in\mathcal{D}_{k}}\left\Vert \boldsymbol{x}_{i}-\boldsymbol{\mu}_{k}\right\Vert _{2}^{2}\)\( \)\(\implies \boldsymbol{x}_{i}\in\mathcal{D}_{s\left(\boldsymbol{x}_{i}\right)} \; \text{where} \; s\left(\boldsymbol{x}_{i}\right)=\arg\min_{k}\left\Vert \boldsymbol{x}_{i}-\boldsymbol{\mu}_{k}\right\Vert _{2}^{2}\)$Step II:
Assume fixed clusters \(\left\{ \mathcal{D}_{k}\right\} \), find the optimal centroids \(\left\{ \boldsymbol{\mu}_{k}\right\} \): $\(\arg\min_{\left\{ \boldsymbol{\mu}_{k}\right\} }\sum_{k=1}^{K}\sum_{\boldsymbol{x}_{i}\in\mathcal{D}_{k}}\left\Vert \boldsymbol{x}_{i}-\boldsymbol{\mu}_{k}\right\Vert _{2}^{2}\)\( \)\(\implies\boldsymbol{\mu}_{k}=\frac{1}{\left|\mathcal{D}_{k}\right|}\sum_{\boldsymbol{x}_{i}\in\mathcal{D}_{k}}\boldsymbol{x}_{i}\)$Step III:
Check for convergence (Change in assignments / location of the center). If not, go to Step I.
(#) The K-Means is implemented in
KMeans.(#) Some implementations of the algorithm supports different metrics.
(?) With regard to the
RGBfeatures, is the metric used is theSquared Euclidean?
The K-Means Algorithm#
Implement the K-Means algorithm as a SciKit Learn compatible class.
(#) The implementation will allow different metrics.
# Implement the K-Means as an Estimator
class KMeansCluster(ClusterMixin, BaseEstimator):
def __init__(self, numClusters: int, numIter: int = 1000, metricType: str = 'sqeuclidean'):
#===========================Fill This===========================#
# 1. Add `numClusters` as an attribute of the object.
# 2. Add `numIter` as an attribute of the object.
# 3. Add `metricType` as an attribute of the object.
# !! The `metricType` must match the values of SciPy's `cdist()`: 'euclidean', 'cityblock', 'seuclidean', 'sqeuclidean', 'cosine', 'correlation'.
self.numClusters = numClusters
self.numIter = numIter
self.metricType = metricType
#===============================================================#
pass
def fit(self, mX: np.ndarray, vY: Optional[np.ndarray] = None) -> Self:
numSamples = mX.shape[0]
featuresDim = mX.shape[1]
if (numSamples < self.numClusters):
raise ValueError(f'The number of samples: {numSamples} should not be smaller than the number of clusters: {self.numClusters}.')
mC = mX[np.random.choice(numSamples, self.numClusters, replace = False)] #<! Centroids (Random initialization)
vL = np.zeros(numSamples, dtype = np.int64) #<! Labels
vF = np.zeros(numSamples, dtype = np.bool_) #<! Flags for each label
#===========================Fill This===========================#
# 1. Create a loop of the number of samples.
# 2. Create the distance matrix between each sample to the centroids (Use the appropriate metrics).
# 3. Extract the labels.
# 4. Iterate on each label group to update the centroids.
# !! You may find `cdist()` from SciPy useful.
# !! Use `mean()` to calculate the centroids.
for ii in range(self.numIter):
mD = sp.spatial.distance.cdist(mX, mC, self.metricType) #<! Distance Matrix (numSamples, numClusters)
vL = np.argmin(mD, axis = 1, out = vL)
for kk in range(numClusters):
# Update `mC`
vF = np.equal(vL, kk, out = vF)
if np.any(vF):
mC[kk, :] = np.mean(mX[vL == kk, :], axis = 0)
#===============================================================#
# SciKit Learn's `KMeans` compatibility
self.cluster_centers_ = mC
self.labels_ = vL
self.inertia_ = np.sum(np.amin(mD, axis = 1))
self.n_iter_ = self.numIter
self.n_features_in = featuresDim
return self
def transform(self, mX):
return sp.spatial.distance.cdist(mX, self.cluster_centers_, self.metricType)
def predict(self, mX):
vL = np.argmin(self.transform(mX), axis = 1)
return vL
def score(self, mX: np.ndarray, vY: Optional[np.ndarray] = None):
# Return the opposite of inertia as the score
mD = self.transform(mX)
inertiaVal = np.sum(np.amin(mD, axis = 1))
return -inertiaVal
(?) Why do the
fit()andpredict()method have thevYinput?(?) If one selects
'cosine'as the distance metric, does the centroid match the metric?(@) Add an option for
K-Means++initialization.(?) How can one use K-Means in an online fashion? Think of a static and dynamic case.
(?) Is the above implementation static or dynamic?
(@) Add a stopping criteria: The maximum movement of a centroid is below a threshold.
Super Pixel Clustering by K-Means#
# Construct the Model & Fit to Data
#===========================Fill This===========================#
# 1. Construct the `KMeansCluster` object.
# 2. Fit it to data.
oKMeans = KMeansCluster(numClusters = numClusters, numIter = numIter)
oKMeans = oKMeans.fit(mX)
#===============================================================#
(?) What’s the difference between
fit()andpredict()in the context of K-Means?
# Extract the Labels and Form a Segmentation Mask
#===========================Fill This===========================#
# 1. Extract the labels of the pixels (Cluster index).
# 2. Reshape them into a mask which matches the image.
mSuperPixel = np.reshape(oKMeans.labels_, (numRows, numCols))
#===============================================================#
# Display Results
hF, hA = plt.subplots(figsize = (12, 8))
PlotSuperPixels(mI, mSuperPixel, hA = hA)
plt.show()
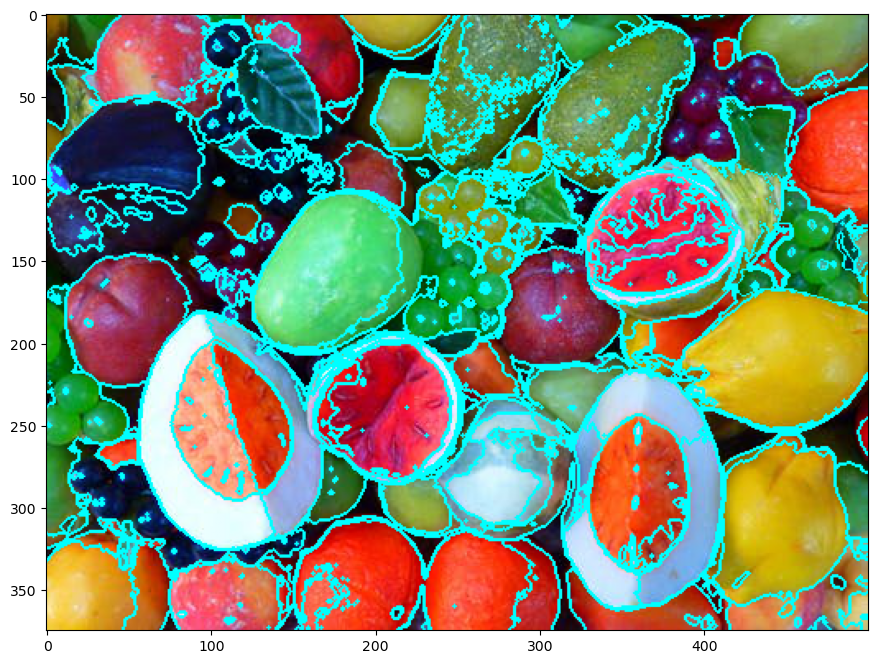

(?) How do we set the hyper parameter
KforK-Means?(?) Why are the separating lines not straight?
(@) Run the notebook again yet using SciKit Learn
KMeansclass. Compare speed.
Analysis of Hyper Parameter K#
One common method to find the optimal K is the Knee / Elbow Method.
In this method a score (Inertia) is plotted vs. the K parameter.
One way to define the elbow point is by the point maximizing the curvature.
This point can be easily calculated by the kneed package.
(#) An alternative to the elbow method is using Silhouette. See Stop Using the Elbow Method, Selecting the Number of Clusters with Silhouette Analysis on KMeans Clustering.
# Score per K
# Takes 4-6 minutes!
lK = [10, 25, 50, 100, 150, 200, 500]
numK = len(lK)
vS = np.full(shape = numK, fill_value = np.nan)
for ii, kVal in enumerate(lK):
oKMeans = KMeansCluster(numClusters = kVal, numIter = numIter)
oKMeans = oKMeans.fit(mX)
vS[ii] = oKMeans.score(mX)
print(f'Finished processing the {ii} / {numK} iteration with K = {kVal}')
Finished processing the 0 / 7 iteration with K = 10
Finished processing the 1 / 7 iteration with K = 25
Finished processing the 2 / 7 iteration with K = 50
Finished processing the 3 / 7 iteration with K = 100
Finished processing the 4 / 7 iteration with K = 150
Finished processing the 5 / 7 iteration with K = 200
Finished processing the 6 / 7 iteration with K = 500
# Plot the Score
hF, hA = plt.subplots(figsize = (12, 8))
hA.plot(lK, vS, label = 'Score')
hA.set_xlabel('Number of Clusters (K)')
hA.set_ylabel('Score (Inertia)')
hA.set_title('Score vs. Number of Clusters')
plt.show()
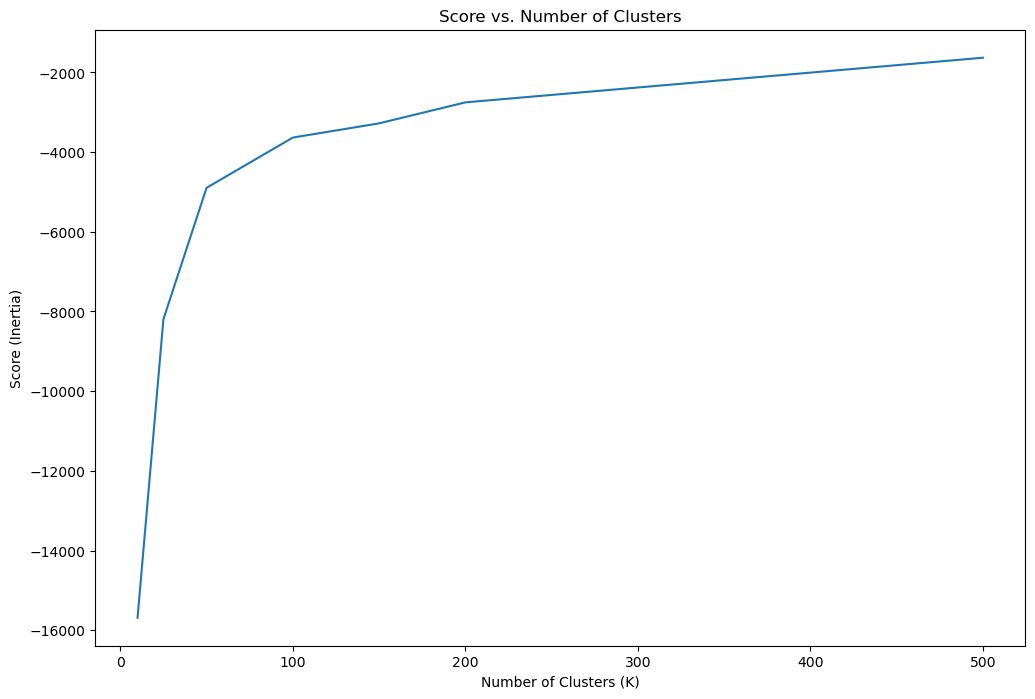
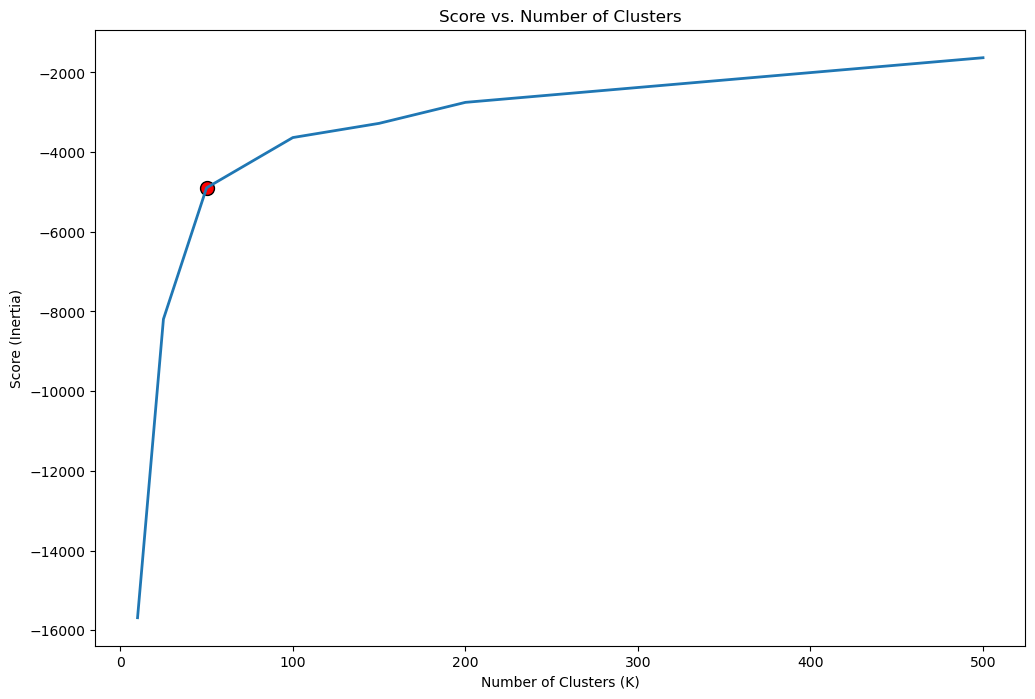
# Locate the Knee Point by Maximum Curvature
oKneeLoc = KneeLocator(lK, vS, curve = 'concave', direction = 'increasing')
kneeK = round(oKneeLoc.knee)
kneeIdx = lK.index(kneeK)
# Plot the Knee
hF, hA = plt.subplots(figsize = (12, 8))
hA.plot(lK, vS, lw = 2, label = 'Score')
hA.scatter(lK[kneeIdx], vS[kneeIdx], s = 100, c = 'r', edgecolors = 'k', label = 'Knee')
hA.set_xlabel('Number of Clusters (K)')
hA.set_ylabel('Score (Inertia)')
hA.set_title('Score vs. Number of Clusters')
plt.show()