Linear Classifier MOONs#
Notebook by:
Royi Avital RoyiAvital@fixelalgorithms.com
Revision History#
Version |
Date |
User |
Content / Changes |
|---|---|---|---|
1.0.000 |
02/03/2024 |
Royi Avital |
First version |
# Import Packages
# General Tools
import numpy as np
import scipy as sp
import pandas as pd
# Machine Learning
from sklearn.datasets import make_moons
# Image Processing
# Machine Learning
# Miscellaneous
import os
from platform import python_version
import random
import timeit
# Typing
from typing import Callable, List, Tuple
# Visualization
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
# from bokeh.plotting import figure, show
# Jupyter
from IPython import get_ipython
from IPython.display import Image, display
from ipywidgets import Dropdown, FloatSlider, interact, IntSlider, Layout
Notations#
(?) Question to answer interactively.
(!) Simple task to add code for the notebook.
(@) Optional / Extra self practice.
(#) Note / Useful resource / Food for thought.
Code Notations:
someVar = 2; #<! Notation for a variable
vVector = np.random.rand(4) #<! Notation for 1D array
mMatrix = np.random.rand(4, 3) #<! Notation for 2D array
tTensor = np.random.rand(4, 3, 2, 3) #<! Notation for nD array (Tensor)
tuTuple = (1, 2, 3) #<! Notation for a tuple
lList = [1, 2, 3] #<! Notation for a list
dDict = {1: 3, 2: 2, 3: 1} #<! Notation for a dictionary
oObj = MyClass() #<! Notation for an object
dfData = pd.DataFrame() #<! Notation for a data frame
dsData = pd.Series() #<! Notation for a series
hObj = plt.Axes() #<! Notation for an object / handler / function handler
Code Exercise#
Single line fill
vallToFill = ???
Multi Line to Fill (At least one)
# You need to start writing
????
Section to Fill
#===========================Fill This===========================#
# 1. Explanation about what to do.
# !! Remarks to follow / take under consideration.
mX = ???
???
#===============================================================#
# Configuration
# %matplotlib inline
seedNum = 512
np.random.seed(seedNum)
random.seed(seedNum)
# Matplotlib default color palette
lMatPltLibclr = ['#1f77b4', '#ff7f0e', '#2ca02c', '#d62728', '#9467bd', '#8c564b', '#e377c2', '#7f7f7f', '#bcbd22', '#17becf']
# sns.set_theme() #>! Apply SeaBorn theme
runInGoogleColab = 'google.colab' in str(get_ipython())
# Constants
FIG_SIZE_DEF = (8, 8)
ELM_SIZE_DEF = 50
CLASS_COLOR = ('b', 'r')
EDGE_COLOR = 'k'
MARKER_SIZE_DEF = 10
LINE_WIDTH_DEF = 2
# Courses Packages
import sys
sys.path.append('../')
sys.path.append('../../')
sys.path.append('../../../')
from utils.DataVisualization import Plot2DLinearClassifier, PlotBinaryClassData
# General Auxiliary Functions
# Parameters
# Data Generation
numSamples = 500
noiseLevel = 0.1
# Data Visualization
numGridPts = 250
Generate / Load Data#
We’ll use the the classic moons data set.
By default it labels the data \({y}_{i} \in \left\{ 0, 1 \right\}\).
We’ll transform it into \({y}_{i} \in \left\{ -1, 1 \right\}\).
# Generate Data
mX, vY = make_moons(n_samples = numSamples, noise = noiseLevel)
print(f'The features data shape: {mX.shape}')
print(f'The labels data shape: {vY.shape}')
The features data shape: (500, 2)
The labels data shape: (500,)
# The Labels
# The labels of the data
print(f'The unique values of the labels: {np.unique(vY)}')
The unique values of the labels: [0 1]
(?) Do the labels fit the model? What should be done?
# Labels Transformation
# Transforming the Labels into {-1, 1}
vY[vY == 0] = -1
# The Labels
# The updated labels
print(f'The unique values of the labels: {np.unique(vY)}')
The unique values of the labels: [-1 1]
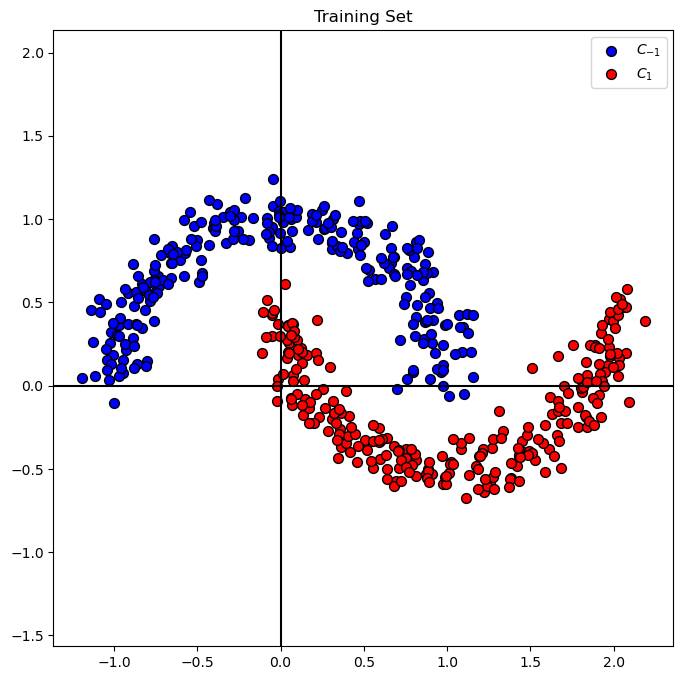
Plot the Data#
# Plot the Data
hA = PlotBinaryClassData(mX, vY, axisTitle = 'Training Set')
Linear Classifier Training#
Training Optimization Problem#
In ideal world, we’d like to optimize:
Where
# Stack the constant column into `mX`
mX = np.column_stack((-np.ones(numSamples), mX))
mX
array([[-1. , 0.75042724, 0.53523666],
[-1. , -0.1674125 , 1.00862292],
[-1. , 0.22313782, 0.87931898],
...,
[-1. , 1.48488659, -0.24967477],
[-1. , 1.59026907, -0.2386105 ],
[-1. , 1.89492566, 0.0257667 ]])
(?) What are the dimensions of
mX?
# The updated dimensions
print(f'The features data shape: {mX.shape}')
The features data shape: (500, 3)
Yet, since the \(\operatorname{sign} \left( \cdot \right)\) isn’t smooth nor continuous we need to approximate it.
The classic candidate is the Sigmoid Function (Member of the S Shaped function family):
See scipy.special.expit() for \(\frac{ 1 }{ 1 + \exp \left( -x \right) }\).
(#) In practice such function requires numerical stable implementation. Use professionally made implementations if available.
The Sigmoid Function derivative is given by:
(#) For derivation of the last step, see https://math.stackexchange.com/questions/78575.
The Loss Function#
Then, using the Sigmoid approximation the loss function becomes (With mean over all data samples \(N\)):
The gradient becomes:
(?) Is the problem convex?
(#) For classification the _Squared \({L}^{2}\) Loss is replaced with Cross Entropy Loss which has better properties in the context of optimization for classification.
(#) For information about the objective function in the context of classification see Stanley Chan - Purdue University - ECE595 / STAT598: Machine Learning I Lecture 14 Logistic Regression.
# Defining the Functions
def SigmoidFun( vX: np.ndarray ):
return (2 * sp.special.expit(vX)) - 1
def GradSigmoidFun(vX: np.ndarray):
vExpit = sp.special.expit(vX)
return 2 * vExpit * (1 - vExpit)
def LossFun(mX: np.ndarray, vW: np.ndarray, vY: np.ndarray):
numSamples = mX.shape[0]
vR = SigmoidFun(mX @ vW) - vY
return np.sum(np.square(vR)) / (4 * numSamples)
def GradLossFun(mX: np.ndarray, vW: np.ndarray, vY: np.ndarray):
numSamples = mX.shape[0]
return (mX.T * GradSigmoidFun(mX @ vW).T) @ (SigmoidFun(mX @ vW) - vY) / (2 * numSamples)
The Gradient Descent#
# Gradient Descent
# Parameters
K = 1000 #<! Num Steps
µ = 0.10 #<! Step Size
vW = np.array([0.0, -1.0, 2.0]) #<! Initial w
mW = np.zeros(shape = (vW.shape[0], K)) #<! Model Parameters (Weights)
vE = np.full(shape = K, fill_value = None) #<! Errors
vL = np.full(shape = K, fill_value = None) #<! Loss
vHatY = np.sign(mX @ vW) #<! Apply the classifier
mW[:, 0] = vW
vE[0] = np.mean(vHatY != vY)
vL[0] = LossFun(mX, vW, vY)
for kk in range(1, K):
vW -= µ * GradLossFun(mX, vW, vY)
mW[:, kk] = vW
vHatY = np.sign(mX @ vW) #<! Apply the classifier
vE[kk] = np.mean(vHatY != vY) #<! Mean Error
vL[kk] = LossFun(mX, vW, vY) #<! Loss Function
# Plotting Function
# Grid of the data support
vV = np.linspace(-2, 2, numGridPts)
mX1, mX2 = np.meshgrid(vV, vV)
def PlotLinClassTrain(itrIdx, mX, mW, vY, K, µ, vE, vL, mX1, mX2):
hF, _ = plt.subplots(nrows = 1, ncols = 2, figsize = (12, 6))
hA1, hA2 = hF.axes[0], hF.axes[1]
# hA1.cla()
# hA2.cla()
Plot2DLinearClassifier(mX, vY, mW[:, itrIdx], mX1, mX2, hA1)
vEE = vE[:itrIdx]
vLL = vL[:itrIdx]
hA2.plot(vEE, color = 'k', lw = 2, label = r'$J \left( w \right)$')
hA2.plot(vLL, color = 'm', lw = 2, label = r'$\tilde{J} \left( w \right)$')
hA2.set_title('Objective Function')
hA2.set_xlabel('Iteration Index')
hA2.set_ylabel('Value')
hA2.set_xlim((0, K - 1))
hA2.set_ylim((0, 1))
hA2.grid()
hA2.legend()
# hF.canvas.draw()
plt.show()
# Display the Optimization Path
# hF, hA = plt.subplots(nrows = 1, ncols = 2, figsize = (12, 6))
# hPlotLinClassTrain = lambda itrIdx: PlotLinClassTrain(itrIdx, mX, mW, vY, K, µ, vE, vL, mX1, mX2, hF)
hPlotLinClassTrain = lambda itrIdx: PlotLinClassTrain(itrIdx, mX[:, 1:], mW, vY, K, µ, vE, vL, mX1, mX2)
kSlider = IntSlider(min = 0, max = K - 1, step = 1, value = 0, layout = Layout(width = '30%'))
interact(hPlotLinClassTrain, itrIdx = kSlider)
# plt.show()
<function __main__.<lambda>(itrIdx)>
(!) Optimize the parameters \(K\) and \(\mu\) to achieve accuracy of ~85% with the least steps.
