Random Forests#
Notebook by:
Royi Avital RoyiAvital@fixelalgorithms.com
Revision History#
Version |
Date |
User |
Content / Changes |
|---|---|---|---|
1.0.000 |
08/04/2024 |
Royi Avital |
First version |
# Import Packages
# General Tools
import numpy as np
import scipy as sp
import pandas as pd
# Machine Learning
from sklearn.datasets import fetch_openml
from sklearn.ensemble import RandomForestClassifier
from sklearn.inspection import permutation_importance
from sklearn.model_selection import train_test_split
# Miscellaneous
import math
import os
from platform import python_version
import random
import timeit
# Typing
from typing import Callable, Dict, List, Optional, Self, Set, Tuple, Union
# Visualization
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
# Jupyter
from IPython import get_ipython
from IPython.display import Image
from IPython.display import display
from ipywidgets import Dropdown, FloatSlider, interact, IntSlider, Layout, SelectionSlider
from ipywidgets import interact
Notations#
(?) Question to answer interactively.
(!) Simple task to add code for the notebook.
(@) Optional / Extra self practice.
(#) Note / Useful resource / Food for thought.
Code Notations:
someVar = 2; #<! Notation for a variable
vVector = np.random.rand(4) #<! Notation for 1D array
mMatrix = np.random.rand(4, 3) #<! Notation for 2D array
tTensor = np.random.rand(4, 3, 2, 3) #<! Notation for nD array (Tensor)
tuTuple = (1, 2, 3) #<! Notation for a tuple
lList = [1, 2, 3] #<! Notation for a list
dDict = {1: 3, 2: 2, 3: 1} #<! Notation for a dictionary
oObj = MyClass() #<! Notation for an object
dfData = pd.DataFrame() #<! Notation for a data frame
dsData = pd.Series() #<! Notation for a series
hObj = plt.Axes() #<! Notation for an object / handler / function handler
Code Exercise#
Single line fill
vallToFill = ???
Multi Line to Fill (At least one)
# You need to start writing
????
Section to Fill
#===========================Fill This===========================#
# 1. Explanation about what to do.
# !! Remarks to follow / take under consideration.
mX = ???
???
#===============================================================#
# Configuration
# %matplotlib inline
seedNum = 512
np.random.seed(seedNum)
random.seed(seedNum)
# Matplotlib default color palette
lMatPltLibclr = ['#1f77b4', '#ff7f0e', '#2ca02c', '#d62728', '#9467bd', '#8c564b', '#e377c2', '#7f7f7f', '#bcbd22', '#17becf']
# sns.set_theme() #>! Apply SeaBorn theme
runInGoogleColab = 'google.colab' in str(get_ipython())
# Constants
FIG_SIZE_DEF = (8, 8)
ELM_SIZE_DEF = 50
CLASS_COLOR = ('b', 'r')
EDGE_COLOR = 'k'
MARKER_SIZE_DEF = 10
LINE_WIDTH_DEF = 2
# Courses Packages
# General Auxiliary Functions
Random Forests Classification#
In this note book we’ll use the Random Forests based classifier in the task of estimating whether a passenger on the Titanic will or will not survive.
We’ll focus on basic pre processing of the data and analyzing the importance of features using the classifier.
(#) This is a very popular data set for classification. You may have a look for notebooks on the need with a deeper analysis of the data set itself.
# Parameters
# Data
trainRatio = 0.75
# Model
numEst = 150
spliCrit = 'gini'
maxLeafNodes = 20
outBagScore = True
# Feature Permutation
numRepeats = 50
Generate / Load Data#
# Load Data
dfX, dsY = fetch_openml('titanic', version = 1, return_X_y = True, as_frame = True, parser = 'auto')
print(f'The features data shape: {dfX.shape}')
print(f'The labels data shape: {dsY.shape}')
The features data shape: (1309, 13)
The labels data shape: (1309,)
Plot Data#
# Plot the Data
# The Data Frame of Features
dfX.head(10)
| pclass | name | sex | age | sibsp | parch | ticket | fare | cabin | embarked | boat | body | home.dest | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 1 | Allen, Miss. Elisabeth Walton | female | 29.0000 | 0 | 0 | 24160 | 211.3375 | B5 | S | 2 | NaN | St Louis, MO |
| 1 | 1 | Allison, Master. Hudson Trevor | male | 0.9167 | 1 | 2 | 113781 | 151.5500 | C22 C26 | S | 11 | NaN | Montreal, PQ / Chesterville, ON |
| 2 | 1 | Allison, Miss. Helen Loraine | female | 2.0000 | 1 | 2 | 113781 | 151.5500 | C22 C26 | S | NaN | NaN | Montreal, PQ / Chesterville, ON |
| 3 | 1 | Allison, Mr. Hudson Joshua Creighton | male | 30.0000 | 1 | 2 | 113781 | 151.5500 | C22 C26 | S | NaN | 135.0 | Montreal, PQ / Chesterville, ON |
| 4 | 1 | Allison, Mrs. Hudson J C (Bessie Waldo Daniels) | female | 25.0000 | 1 | 2 | 113781 | 151.5500 | C22 C26 | S | NaN | NaN | Montreal, PQ / Chesterville, ON |
| 5 | 1 | Anderson, Mr. Harry | male | 48.0000 | 0 | 0 | 19952 | 26.5500 | E12 | S | 3 | NaN | New York, NY |
| 6 | 1 | Andrews, Miss. Kornelia Theodosia | female | 63.0000 | 1 | 0 | 13502 | 77.9583 | D7 | S | 10 | NaN | Hudson, NY |
| 7 | 1 | Andrews, Mr. Thomas Jr | male | 39.0000 | 0 | 0 | 112050 | 0.0000 | A36 | S | NaN | NaN | Belfast, NI |
| 8 | 1 | Appleton, Mrs. Edward Dale (Charlotte Lamson) | female | 53.0000 | 2 | 0 | 11769 | 51.4792 | C101 | S | D | NaN | Bayside, Queens, NY |
| 9 | 1 | Artagaveytia, Mr. Ramon | male | 71.0000 | 0 | 0 | PC 17609 | 49.5042 | NaN | C | NaN | 22.0 | Montevideo, Uruguay |
(?) What kind of a feature is
name? Should it be used?(?) What kind of a feature is
ticket? Should it be used?
Pre Processing and EDA#
# Dropping Features
# Dropping the `name` and `ticket` features (Avoid 1:1 identification)
dfX = dfX.drop(columns = ['name', 'ticket'])
(#) Pay attention that we dropped the features, but a deeper analysis could extract information from them (Type of ticket, families, etc…).
# The Features Data Frame
dfX.head(10)
| pclass | sex | age | sibsp | parch | fare | cabin | embarked | boat | body | home.dest | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 1 | female | 29.0000 | 0 | 0 | 211.3375 | B5 | S | 2 | NaN | St Louis, MO |
| 1 | 1 | male | 0.9167 | 1 | 2 | 151.5500 | C22 C26 | S | 11 | NaN | Montreal, PQ / Chesterville, ON |
| 2 | 1 | female | 2.0000 | 1 | 2 | 151.5500 | C22 C26 | S | NaN | NaN | Montreal, PQ / Chesterville, ON |
| 3 | 1 | male | 30.0000 | 1 | 2 | 151.5500 | C22 C26 | S | NaN | 135.0 | Montreal, PQ / Chesterville, ON |
| 4 | 1 | female | 25.0000 | 1 | 2 | 151.5500 | C22 C26 | S | NaN | NaN | Montreal, PQ / Chesterville, ON |
| 5 | 1 | male | 48.0000 | 0 | 0 | 26.5500 | E12 | S | 3 | NaN | New York, NY |
| 6 | 1 | female | 63.0000 | 1 | 0 | 77.9583 | D7 | S | 10 | NaN | Hudson, NY |
| 7 | 1 | male | 39.0000 | 0 | 0 | 0.0000 | A36 | S | NaN | NaN | Belfast, NI |
| 8 | 1 | female | 53.0000 | 2 | 0 | 51.4792 | C101 | S | D | NaN | Bayside, Queens, NY |
| 9 | 1 | male | 71.0000 | 0 | 0 | 49.5042 | NaN | C | NaN | 22.0 | Montevideo, Uruguay |
# The Labels Data Series
dsY.head(10)
0 1
1 1
2 0
3 0
4 0
5 1
6 1
7 0
8 1
9 0
Name: survived, dtype: category
Categories (2, object): ['0', '1']
# Merging Data
dfData = pd.concat([dfX, dsY], axis = 1)
# Data Frame Info
dfData.info()
<class 'pandas.core.frame.DataFrame'>
RangeIndex: 1309 entries, 0 to 1308
Data columns (total 12 columns):
# Column Non-Null Count Dtype
--- ------ -------------- -----
0 pclass 1309 non-null int64
1 sex 1309 non-null category
2 age 1046 non-null float64
3 sibsp 1309 non-null int64
4 parch 1309 non-null int64
5 fare 1308 non-null float64
6 cabin 295 non-null object
7 embarked 1307 non-null category
8 boat 486 non-null object
9 body 121 non-null float64
10 home.dest 745 non-null object
11 survived 1309 non-null category
dtypes: category(3), float64(3), int64(3), object(3)
memory usage: 96.4+ KB
# Missing / Invalid Values
# Null / NA / NaN Matrix
#===========================Fill This===========================#
# 1. Calculate the logical map of invalid values using the method `isna()`.
mInvData = dfData.isna() #<! The logical matrix of invalid values
#===============================================================#
hF, hA = plt.subplots(figsize = FIG_SIZE_DEF)
sns.heatmap(data = mInvData, square = False, ax = hA)
hA.set_title('Invalid Data Map')
plt.show()
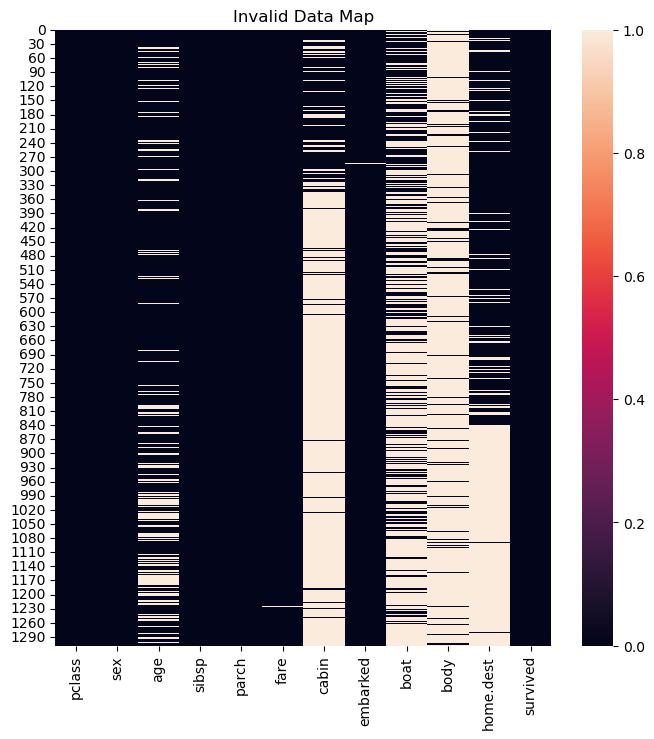
Since the features cabin, boat, body and home.dest have mostly non valid values the will be dropped as well.
(#) Some implementation of Ensemble Trees can handle missing values. They might benefit in such case asl well.
# Features Filtering
# Removing Features with Invalid Values.
dfData = dfData.drop(columns = ['cabin', 'boat', 'body', 'home.dest'])
# Data Visualization
# Basic EDA on the Data
numCol = dfData.shape[1]
lCols = dfData.columns
numAx = int(np.ceil(np.sqrt(numCol)))
# hIsCatLikData = lambda dsX: (pd.api.types.is_categorical_dtype(dsX) or pd.api.types.is_bool_dtype(dsX) or pd.api.types.is_object_dtype(dsX) or pd.api.types.is_integer_dtype(dsX)) #<! Deprecated
hIsCatLikData = lambda dsX: (isinstance(dsX.dtype, pd.CategoricalDtype) or pd.api.types.is_bool_dtype(dsX) or pd.api.types.is_object_dtype(dsX) or pd.api.types.is_integer_dtype(dsX))
hF, hAs = plt.subplots(nrows = numAx, ncols = numAx, figsize = (16, 12))
hAs = hAs.flat
for ii in range(numCol):
colName = dfData.columns[ii]
if hIsCatLikData(dfData[colName]):
sns.histplot(data = dfData, x = colName, hue = 'survived', stat = 'count', discrete = True, common_norm = True, multiple = 'dodge', ax = hAs[ii])
else:
sns.kdeplot(data = dfData, x = colName, hue = 'survived', fill = True, common_norm = True, ax = hAs[ii])
plt.show()
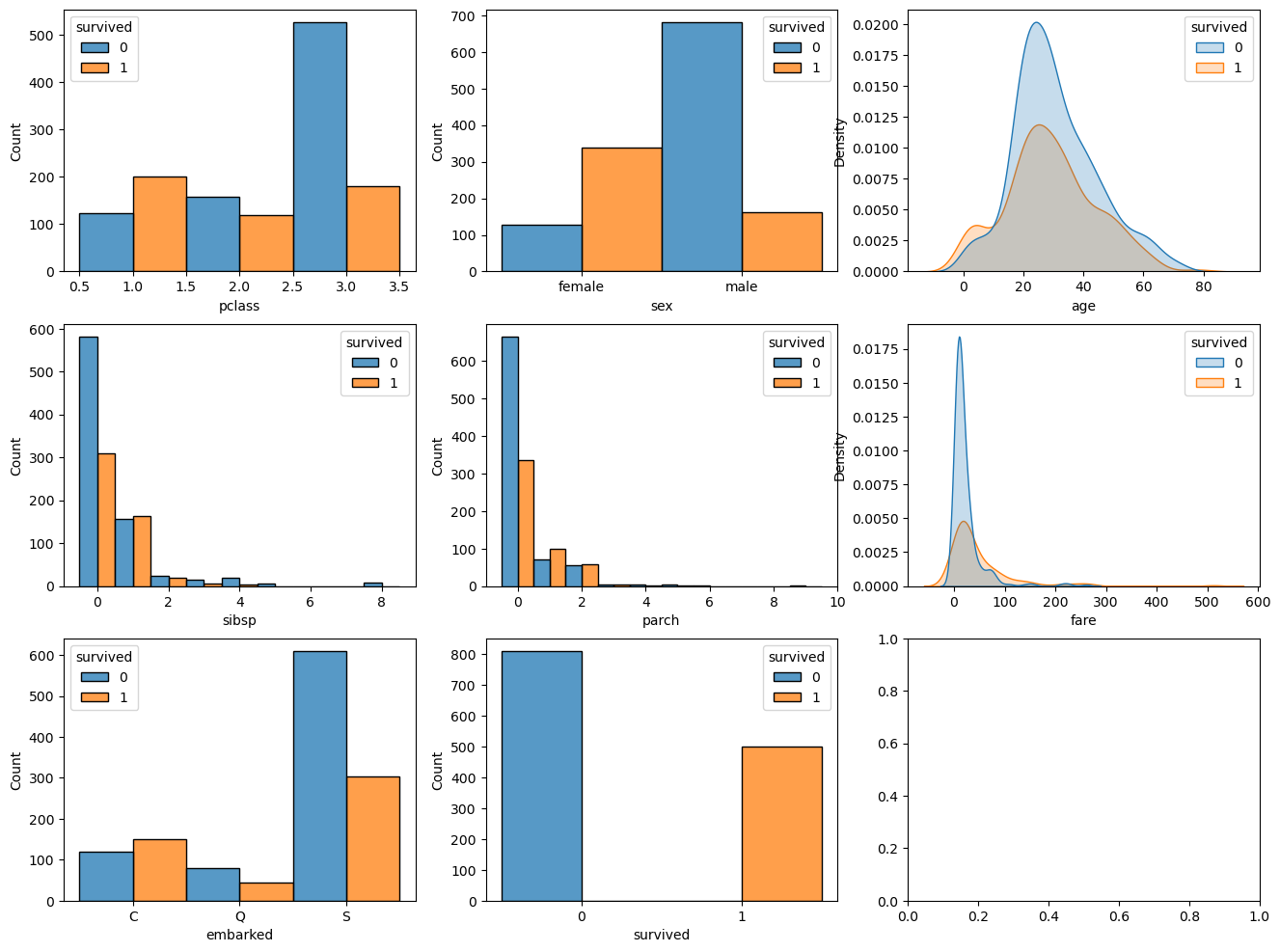
(?) Have a look on the features and try to estimate their importance for the estimation.
Filling Missing Data#
For the rest of the missing values we’ll use a simple method of interpolation:
Categorical Data: Using the mode value.
Numeric Data: Using the median / mean.
(#) We could employ much more delicate and sophisticated data.
For instance, use the mean value of the samepclass. Namely profiling the data by other features to interpolate the missing feature.(#) Data imputing can be done by using a model as well: Regression for continuous data, Classification for categorical data.
(#) The relevant classed in SciKti Learn:
SimpleImputer,IterativeImputer,KNNImputer.
# Missing Value by Dictionary
dNaNs = {'embarked': dfData['embarked'].mode()[0], 'age': dfData['age'].median(), 'fare': dfData['fare'].median()}
dfData = dfData.fillna(value = dNaNs, inplace = False) #<! We can use the `inplace` for efficiency
(#) The above is equivalent of using
SimpleImputer.
# Null / NA / NaN Matrix
mInvData = dfData.isna() #<! The logical matrix of invalid values
hF, hA = plt.subplots(figsize = FIG_SIZE_DEF)
sns.heatmap(data = mInvData, square = False, ax = hA)
hA.set_title('Invalid Data Map')
plt.show()
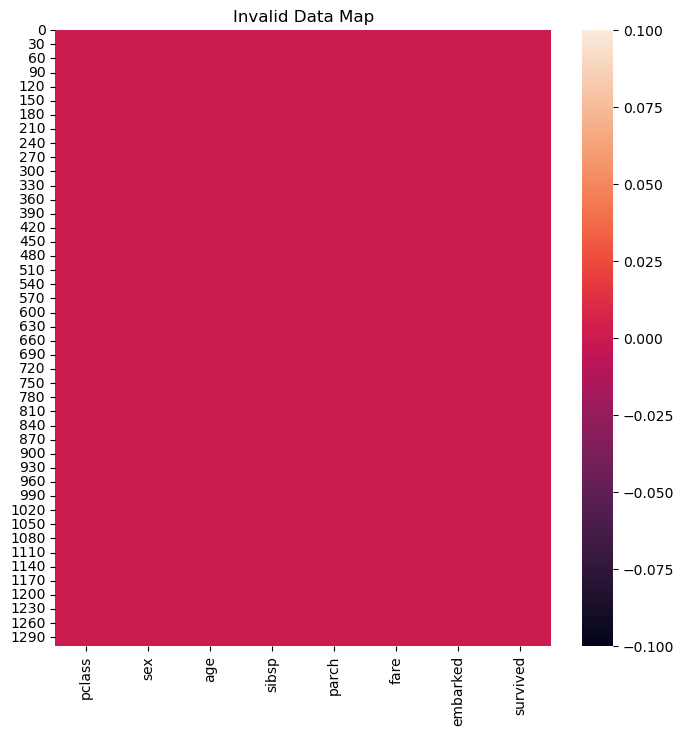
Conversion of Categorical Data#
In this notebook we’ll use the RandomForestClassifier implementation of Random Forests.
At the moment, it doesn’t support Categorical Data, hence we’ll use Dummy Variables (One Hot Encoding).
(#) Pay attention that one hot encoding is an inferior approach to having a real support for categorical data.
(#) For 2 values categorical data (Binary feature) there is no need fo any special support.
(#) Currently, the implementation of
DecisionTreeClassifierdoesn’t support categorical values which are not ordinal (As it treats them as numeric). Hence we must useOneHotEncoder.
See status at scikit-learn/scikit-learn#12866.(#) The One Hot Encoding is not perfect. See Are Categorical Variables Getting Lost in Your Random Forests (Also Notebook - Are Categorical Variables Getting Lost in Your Random Forests).
# Features Encoding
# 1. The feature 'embarked' -> One Hot Encoding.
# 1. The feature 'sex' -> Mapping (Female -> 0, Male -> 1).
dfData = pd.get_dummies(dfData, columns = ['embarked'], drop_first = False)
dfData['sex'] = dfData['sex'].map({'female': 0, 'male': 1})
# Convert Data Type
dfData = dfData.astype(dtype = {'pclass': np.uint8, 'sex': np.uint8, 'sibsp': np.uint8, 'parch': np.uint8, 'survived': np.uint8})
dfData.info()
<class 'pandas.core.frame.DataFrame'>
RangeIndex: 1309 entries, 0 to 1308
Data columns (total 10 columns):
# Column Non-Null Count Dtype
--- ------ -------------- -----
0 pclass 1309 non-null uint8
1 sex 1309 non-null uint8
2 age 1309 non-null float64
3 sibsp 1309 non-null uint8
4 parch 1309 non-null uint8
5 fare 1309 non-null float64
6 survived 1309 non-null uint8
7 embarked_C 1309 non-null bool
8 embarked_Q 1309 non-null bool
9 embarked_S 1309 non-null bool
dtypes: bool(3), float64(2), uint8(5)
memory usage: 30.8 KB
Train the Random Forests Model#
The random forests models basically creates weak classifiers by limiting their access to the data and features.
This basically also limits their correlation which means we can use their mean in order to reduce the variance of the estimation.
Split Train & Test#
# Split to Features and Labels
dfX = dfData.drop(columns = ['survived'])
dsY = dfData['survived']
print(f'The features data shape: {dfX.shape}')
print(f'The labels data shape: {dsY.shape}')
The features data shape: (1309, 9)
The labels data shape: (1309,)
# Split the Data
dfXTrain, dfXTest, dsYTrain, dsYTest = train_test_split(dfX, dsY, train_size = trainRatio, random_state = seedNum, shuffle = True, stratify = dsY)
Train the Model#
# Construct the Model & Train
oRndForestsCls = RandomForestClassifier(n_estimators = numEst, criterion = spliCrit, max_leaf_nodes = maxLeafNodes, oob_score = outBagScore)
oRndForestsCls = oRndForestsCls.fit(dfXTrain, dsYTrain)
# Scores of the Model
print(f'The train accuracy : {oRndForestsCls.score(dfXTrain, dsYTrain):0.2%}')
print(f'The out of bag accuracy: {oRndForestsCls.oob_score_:0.2%}')
print(f'The test accuracy : {oRndForestsCls.score(dfXTest, dsYTest):0.2%}')
The train accuracy : 86.03%
The out of bag accuracy: 83.18%
The test accuracy : 75.30%
(!) Try different values for the model’s hyper parameter (Try defaults as well).
Feature Importance Analysis#
Contribution to Impurity Decrease#
This is the default method for feature importance of decision trees based methods.
It basically sums the amount of impurity reduced by splits by each feature.
# Extract Feature Importance
vFeatImp = oRndForestsCls.feature_importances_
vIdxSort = np.argsort(vFeatImp)
# Display Feature Importance
hF, hA = plt.subplots(figsize = (16, 8))
hA.bar(x = dfXTrain.columns[vIdxSort], height = vFeatImp[vIdxSort])
hA.set_title('Features Importance of the Model')
hA.set_xlabel('Feature Name')
hA.set_title('Importance')
plt.show()
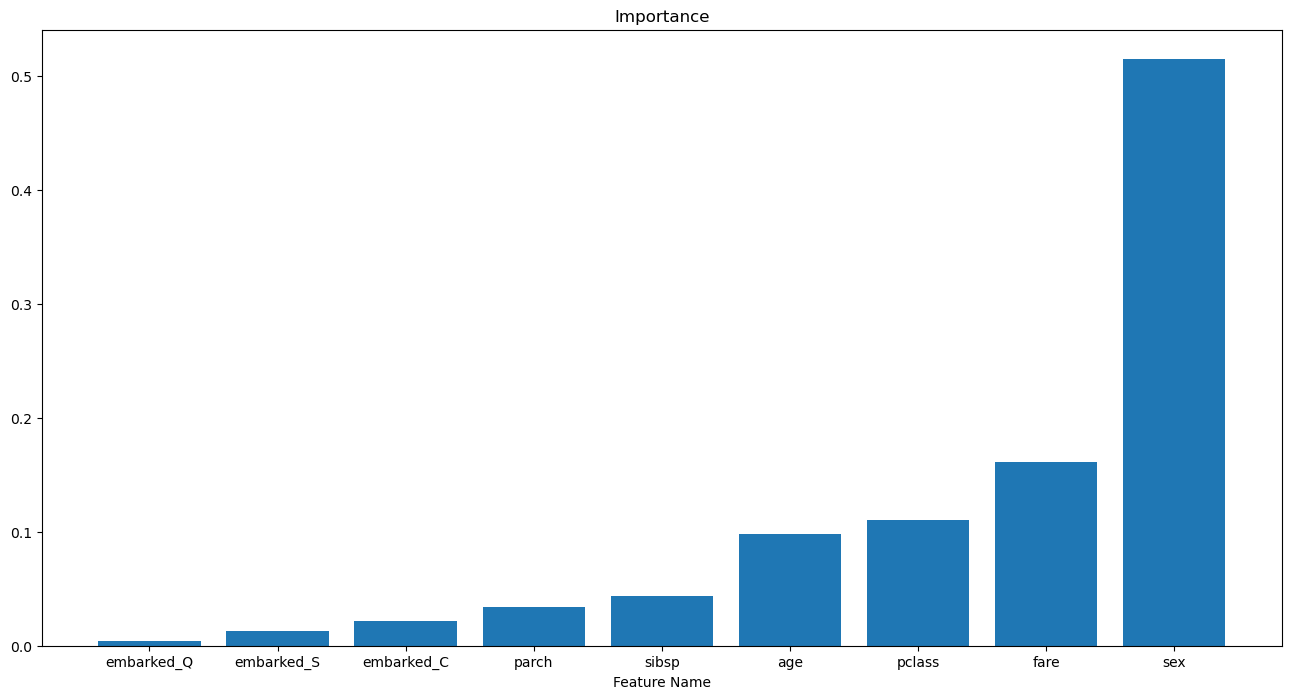
(?) Do we need all 3:
embarked_C,embarked_Qandembarked_S? Look at the options ofget_dummies().
Permutation Effect#
This is a more general method to measure the importance of a feature.
We basically replace the values with “noise” to see how much performance has been deteriorated.
(#) The SciKit Learn’s function which automates the process is
permutation_importance().(#) This is highly time consuming operation. Hence the speed of decision trees based methods creates a synergy.
(#) The importance is strongly linked to the estimator in use.
# Test Permutations
oFeatImpPermTrain = permutation_importance(oRndForestsCls, dfXTrain, dsYTrain, n_repeats = numRepeats)
oFeatImpPermTest = permutation_importance(oRndForestsCls, dfXTest, dsYTest, n_repeats = numRepeats)
# Generate Data Frame
dT = {'Feature': [], 'Importance': [], 'Data': []}
for (dataName, mScore) in [('Train', oFeatImpPermTrain.importances), ('Test', oFeatImpPermTest.importances)]:
for ii, featName in enumerate(dfX.columns):
for jj in range(numRepeats):
dT['Feature'].append(featName)
dT['Importance'].append(mScore[ii, jj])
dT['Data'].append(dataName)
dfFeatImpPerm = pd.DataFrame(dT)
dfFeatImpPerm
| Feature | Importance | Data | |
|---|---|---|---|
| 0 | pclass | 0.078491 | Train |
| 1 | pclass | 0.075433 | Train |
| 2 | pclass | 0.076453 | Train |
| 3 | pclass | 0.089704 | Train |
| 4 | pclass | 0.082569 | Train |
| ... | ... | ... | ... |
| 895 | embarked_S | 0.021341 | Test |
| 896 | embarked_S | 0.018293 | Test |
| 897 | embarked_S | 0.018293 | Test |
| 898 | embarked_S | 0.015244 | Test |
| 899 | embarked_S | 0.018293 | Test |
900 rows × 3 columns
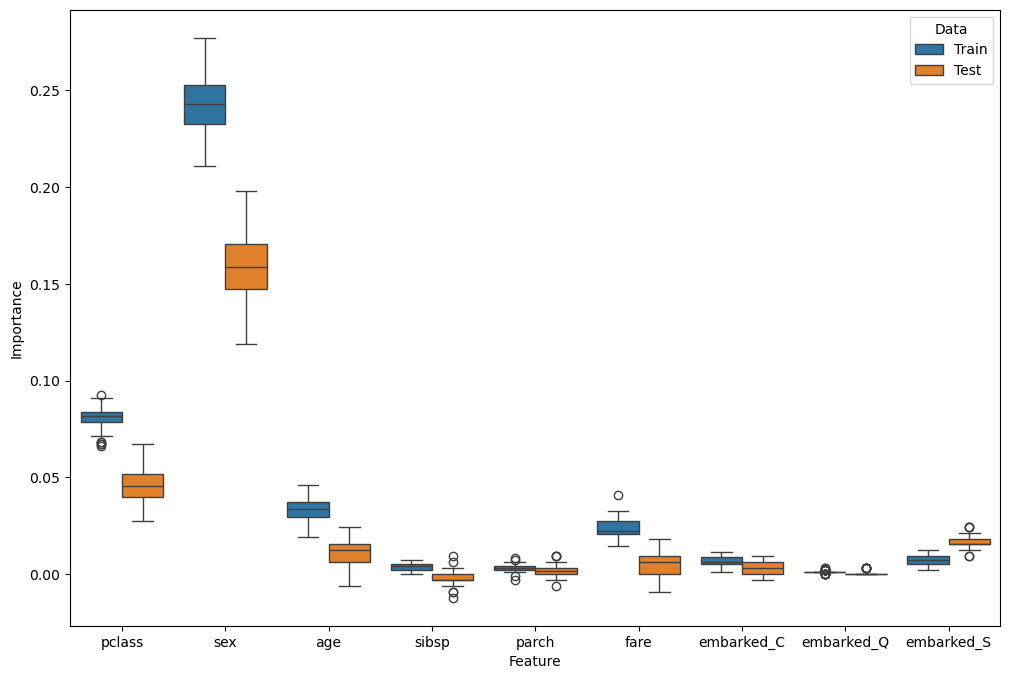
# Plot Results
hF, hA = plt.subplots(figsize = (12, 8))
sns.boxplot(data = dfFeatImpPerm, x = 'Feature', y = 'Importance', hue = 'Data', ax = hA)
plt.show()
(@) Try extracting better results by using the dropped features.
