fashion-mnist#
Notebook by:
Royi Avital RoyiAvital@fixelalgorithms.com
Revision History#
Version |
Date |
User |
Content / Changes |
|---|---|---|---|
1.0.000 |
17/03/2024 |
Royi Avital |
First version |
# Import Packages
# General Tools
import numpy as np
import scipy as sp
import pandas as pd
# Machine Learning
from sklearn.model_selection import GridSearchCV
from sklearn.svm import SVC
# Image Processing
# Machine Learning
# Miscellaneous
import os
from platform import python_version
import random
# Typing
from typing import Callable, Dict, List, Optional, Set, Tuple, Union
# Visualization
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
# Jupyter
from IPython import get_ipython
Notations#
(?) Question to answer interactively.
(!) Simple task to add code for the notebook.
(@) Optional / Extra self practice.
(#) Note / Useful resource / Food for thought.
Code Notations:
someVar = 2; #<! Notation for a variable
vVector = np.random.rand(4) #<! Notation for 1D array
mMatrix = np.random.rand(4, 3) #<! Notation for 2D array
tTensor = np.random.rand(4, 3, 2, 3) #<! Notation for nD array (Tensor)
tuTuple = (1, 2, 3) #<! Notation for a tuple
lList = [1, 2, 3] #<! Notation for a list
dDict = {1: 3, 2: 2, 3: 1} #<! Notation for a dictionary
oObj = MyClass() #<! Notation for an object
dfData = pd.DataFrame() #<! Notation for a data frame
dsData = pd.Series() #<! Notation for a series
hObj = plt.Axes() #<! Notation for an object / handler / function handler
Code Exercise#
Single line fill
vallToFill = ???
Multi Line to Fill (At least one)
# You need to start writing
????
Section to Fill
#===========================Fill This===========================#
# 1. Explanation about what to do.
# !! Remarks to follow / take under consideration.
mX = ???
???
#===============================================================#
# Configuration
# %matplotlib inline
seedNum = 512
np.random.seed(seedNum)
random.seed(seedNum)
# Matplotlib default color palette
lMatPltLibclr = ['#1f77b4', '#ff7f0e', '#2ca02c', '#d62728', '#9467bd', '#8c564b', '#e377c2', '#7f7f7f', '#bcbd22', '#17becf']
# sns.set_theme() #>! Apply SeaBorn theme
runInGoogleColab = 'google.colab' in str(get_ipython())
# Constants
FIG_SIZE_DEF = (8, 8)
ELM_SIZE_DEF = 50
CLASS_COLOR = ('b', 'r')
EDGE_COLOR = 'k'
MARKER_SIZE_DEF = 10
LINE_WIDTH_DEF = 2
# Fashion MNIST
TRAIN_DATA_SET_IMG_URL = r'https://github.com/zalandoresearch/fashion-mnist/raw/master/data/fashion/train-images-idx3-ubyte.gz'
TRAIN_DATA_SET_LBL_URL = r'https://github.com/zalandoresearch/fashion-mnist/raw/master/data/fashion/train-labels-idx1-ubyte.gz'
TEST_DATA_SET_IMG_URL = r'https://github.com/zalandoresearch/fashion-mnist/raw/master/data/fashion/t10k-images-idx3-ubyte.gz'
TEST_DATA_SET_LBL_URL = r'https://github.com/zalandoresearch/fashion-mnist/raw/master/data/fashion/t10k-labels-idx1-ubyte.gz'
TRAIN_DATA_IMG_FILE_NAME = 'TrainImgFile'
TRAIN_DATA_LBL_FILE_NAME = 'TrainLblFile'
TEST_DATA_IMG_FILE_NAME = 'TestImgFile'
TEST_DATA_LBL_FILE_NAME = 'TestLblFile'
TRAIN_DATA_SET_FILE_NAME = 'FashionMnistTrainDataSet.npz'
TEST_DATA_SET_FILE_NAME = 'FashionMnistTestDataSet.npz'
TRAIN_DATA_NUM_IMG = 60_000
TEST_DATA_NUM_IMG = 10_000
D_CLASSES = {0: 'T-Shirt', 1: 'Trouser', 2: 'Pullover', 3: 'Dress', 4: 'Coat', 5: 'Sandal', 6: 'Shirt', 7: 'Sneaker', 8: 'Bag', 9: 'Boots'}
L_CLASSES = ['T-Shirt', 'Trouser', 'Pullover', 'Dress', 'Coat', 'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Boots']
# Courses Packages
import sys,os
sys.path.append('../')
sys.path.append('../../')
sys.path.append('../../../')
from utils.DataVisualization import PlotConfusionMatrix, PlotLabelsHistogram, PlotMnistImages
from utils.DataManipulation import DownloadDecompressGzip, ConvertMnistDataDf
# General Auxiliary Functions
def IsStrFloat(inStr: any) -> bool:
#Support None input
if inStr is None:
return False
try:
float(inStr)
return True
except ValueError:
return False
Model Parameter Optimization with Kernel SVM#
In this exercise we’ll apply the Cross Validation automatically to find the optimal hyper parameters for the Kernel SVM Model.
In order to achieve this we’ll do a Grid Search for Hyper Parameters Optimization.
Load the Fashion MNIST Data Set manually (Done by the notebook).
Train a baseline Linear SVM model.
Find the optimal Kernel SVM model using Grid Search.
Extract the optimal model.
Plot the Confusion Matrix of the best model on the training data.
(#) You may and should use the functions in the
Auxiliary Functionssection.
# Parameters
numSamplesTrain = 4_000
numSamplesTest = 1_000
numImg = 3
# Linear SVM (Baseline Model)
paramC = 1
kernelType = 'linear'
#===========================Fill This===========================#
# 1. Think of the parameters to optimize.
# 2. Select the set to optimize over.
# 3. Set the number of folds in the cross validation.
numFold = 5
#===============================================================#
Generate / Load Data#
Load the Fashion MNIST Data Set.
# Load Data
if os.path.isfile(TRAIN_DATA_SET_FILE_NAME):
dData = np.load(TRAIN_DATA_SET_FILE_NAME)
mXTrain, vYTrain = dData['mXTrain'], dData['vYTrain']
else:
if not os.path.isfile(TRAIN_DATA_IMG_FILE_NAME):
DownloadDecompressGzip(TRAIN_DATA_SET_IMG_URL, TRAIN_DATA_IMG_FILE_NAME) #<! Download Data (GZip File)
if not os.path.isfile(TRAIN_DATA_LBL_FILE_NAME):
DownloadDecompressGzip(TRAIN_DATA_SET_LBL_URL, TRAIN_DATA_LBL_FILE_NAME) #<! Download Data (GZip File)
mXTrain, vYTrain = ConvertMnistDataDf(TRAIN_DATA_IMG_FILE_NAME, TRAIN_DATA_LBL_FILE_NAME)
np.savez_compressed(TRAIN_DATA_SET_FILE_NAME, mXTrain = mXTrain, vYTrain = vYTrain)
if os.path.isfile(TRAIN_DATA_IMG_FILE_NAME):
os.remove(TRAIN_DATA_IMG_FILE_NAME)
if os.path.isfile(TRAIN_DATA_LBL_FILE_NAME):
os.remove(TRAIN_DATA_LBL_FILE_NAME)
if os.path.isfile(TEST_DATA_SET_FILE_NAME):
dData = np.load(TEST_DATA_SET_FILE_NAME)
mXTest, vYTest = dData['mXTest'], dData['vYTest']
else:
if not os.path.isfile(TEST_DATA_IMG_FILE_NAME):
DownloadDecompressGzip(TEST_DATA_SET_IMG_URL, TEST_DATA_IMG_FILE_NAME) #<! Download Data (GZip File)
if not os.path.isfile(TEST_DATA_LBL_FILE_NAME):
DownloadDecompressGzip(TEST_DATA_SET_LBL_URL, TEST_DATA_LBL_FILE_NAME) #<! Download Data (GZip File)
mXTest, vYTest = ConvertMnistDataDf(TEST_DATA_IMG_FILE_NAME, TEST_DATA_LBL_FILE_NAME)
np.savez_compressed(TEST_DATA_SET_FILE_NAME, mXTest = mXTest, vYTest = vYTest)
if os.path.isfile(TEST_DATA_IMG_FILE_NAME):
os.remove(TEST_DATA_IMG_FILE_NAME)
if os.path.isfile(TEST_DATA_LBL_FILE_NAME):
os.remove(TEST_DATA_LBL_FILE_NAME)
vSampleIdx = np.random.choice(mXTrain.shape[0], numSamplesTrain)
mXTrain = mXTrain[vSampleIdx, :]
vYTrain = vYTrain[vSampleIdx]
vSampleIdx = np.random.choice(mXTest.shape[0], numSamplesTest)
mXTest = mXTest[vSampleIdx, :]
vYTest = vYTest[vSampleIdx]
print(f'The number of train data samples: {mXTrain.shape[0]}')
print(f'The number of train features per sample: {mXTrain.shape[1]}')
print(f'The unique values of the train labels: {np.unique(vYTrain)}')
print(f'The number of test data samples: {mXTest.shape[0]}')
print(f'The number of test features per sample: {mXTest.shape[1]}')
print(f'The unique values of the test labels: {np.unique(vYTest)}')
The number of train data samples: 4000
The number of train features per sample: 784
The unique values of the train labels: [0 1 2 3 4 5 6 7 8 9]
The number of test data samples: 1000
The number of test features per sample: 784
The unique values of the test labels: [0 1 2 3 4 5 6 7 8 9]
Pre Process Data#
The image data is in the UInt8 data form with values in {0, 1, 2, ..., 255}.
The pre process step scales it into [0, 1] range.
# Pre Process Data
#===========================Fill This===========================#
# 1. Scale data into [0, 1] range.
mXTrain = mXTrain / 255
mXTest = mXTest / 255
#===============================================================#
### TODO - check if will be the same for each pixel to be 0 or 1 !!!!
Plot Data#
# Plot the Data
hF = PlotMnistImages(mXTrain, vYTrain, numImg, lClasses = L_CLASSES)

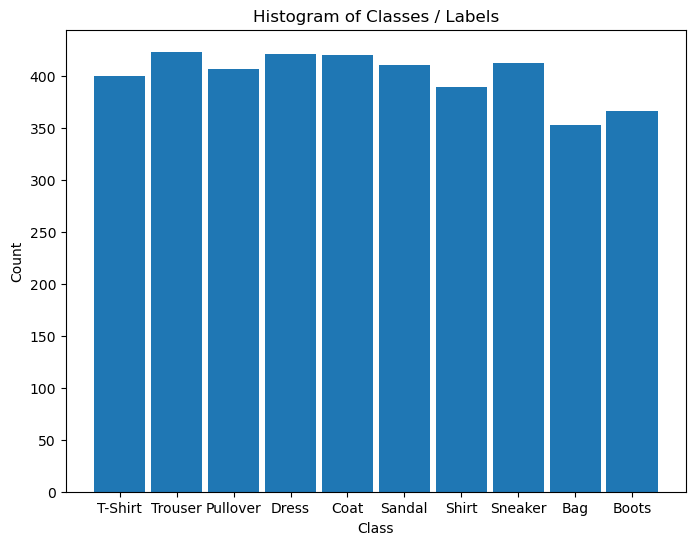
# Plot the Histogram of Classes
hA = PlotLabelsHistogram(vYTrain)
hA.set_xticks(range(len(L_CLASSES)))
hA.set_xticklabels(L_CLASSES)
plt.show()
Train Linear SVM Classifier#
The Linear SVM will function as the baseline classifier.
# SVM Linear Model
#===========================Fill This===========================#
# 1. Construct a baseline model (Linear SVM).
# 2. Train the model.
# 3. Score the model (Accuracy). Keep result in a variable named `modelScore`.
svm_linear = SVC(C = paramC, kernel = kernelType).fit(mXTrain, vYTrain)
modelScore = svm_linear.score(mXTest, vYTest)
#===============================================================#
print(f'The model score (Accuracy) on the data: {modelScore:0.2%}') #<! Accuracy
The model score (Accuracy) on the data: 79.30%
Train Kernel SVM#
In this section we’ll train a Kernel SVM. We’ll find the optimal kernel and other hyper parameters by cross validation.
In order to optimize on the following parameters: C, kernel and gamma we’ll use GridSearchCV().
The idea is iterating over the grid of parameters of the model to find the optimal one.
Each parameterized model is evaluated by a Cross Validation.
In order to use it we need to define:
The Model (
estimator) - Which model is used.The Parameters Grid (
param_grid) - The set of parameter to try.The Scoring (
scoring) - The score used to define the best model.The Cross Validation Iterator (
cv) - The iteration to validate the model.
(#) Pay attention to the expected run time. Using
verboseis useful.(#) This is a classic grid search which is not the most efficient policy. There are more advanced policies.
(#) The
GridSearchCV()is limited to one instance of an estimator.
Yet using Pipelines we may test different types of estimators.
# Construct the Grid Search object
#===========================Fill This===========================#
# 1. Set the parameters to iterate over and their values.
dParams = {
'C' : [1,2,3,4],
'kernel' : ['linear', 'poly', 'rbf', 'sigmoid'],
'gamma' : ["scale", "auto"],
}
#===============================================================#
oGsSvc = GridSearchCV(estimator = SVC(), param_grid = dParams, scoring = None, cv = numFold, verbose = 4,n_jobs=4)
TODO

see also using ParameterGrid#
from sklearn.model_selection import ParameterGrid
lParamGrid = list(ParameterGrid(dParams))
print(f'The number of parameter combinations: {len(lParamGrid)}')
lParamGrid[2]
The number of parameter combinations: 32
{'C': 1, 'gamma': 'scale', 'kernel': 'rbf'}
(#) You may want to have a look at the
n_jobsparameter.
run#
# Hyper Parameter Optimization
# Training the model with each combination of hyper parameters.
#===========================Fill This===========================#
# 1. The model trains on the train data using Stratified K Fold cross validation.
oGsSvc = oGsSvc.fit(mXTrain, vYTrain) #<! It may take few minutes
#===============================================================#
Fitting 5 folds for each of 32 candidates, totalling 160 fits
[CV 1/5] END ...C=1, gamma=scale, kernel=linear;, score=0.820 total time= 5.4s
[CV 3/5] END ...C=1, gamma=scale, kernel=linear;, score=0.816 total time= 5.4s
[CV 4/5] END ...C=1, gamma=scale, kernel=linear;, score=0.823 total time= 5.6s
[CV 2/5] END ...C=1, gamma=scale, kernel=linear;, score=0.841 total time= 5.7s
[CV 5/5] END ...C=1, gamma=scale, kernel=linear;, score=0.828 total time= 2.5s
[CV 1/5] END .....C=1, gamma=scale, kernel=poly;, score=0.787 total time= 2.8s
[CV 2/5] END .....C=1, gamma=scale, kernel=poly;, score=0.794 total time= 2.6s
[CV 3/5] END .....C=1, gamma=scale, kernel=poly;, score=0.779 total time= 4.1s
[CV 5/5] END .....C=1, gamma=scale, kernel=poly;, score=0.807 total time= 2.2s
[CV 4/5] END .....C=1, gamma=scale, kernel=poly;, score=0.786 total time= 3.2s
[CV 1/5] END ......C=1, gamma=scale, kernel=rbf;, score=0.841 total time= 3.1s
[CV 2/5] END ......C=1, gamma=scale, kernel=rbf;, score=0.849 total time= 3.7s
[CV 3/5] END ......C=1, gamma=scale, kernel=rbf;, score=0.816 total time= 3.3s
[CV 4/5] END ......C=1, gamma=scale, kernel=rbf;, score=0.840 total time= 3.3s
[CV 5/5] END ......C=1, gamma=scale, kernel=rbf;, score=0.849 total time= 3.5s
[CV 2/5] END ..C=1, gamma=scale, kernel=sigmoid;, score=0.417 total time= 3.6s
[CV 4/5] END ..C=1, gamma=scale, kernel=sigmoid;, score=0.451 total time= 4.1s
[CV 1/5] END ..C=1, gamma=scale, kernel=sigmoid;, score=0.411 total time= 6.0s
[CV 3/5] END ..C=1, gamma=scale, kernel=sigmoid;, score=0.425 total time= 5.1s
[CV 5/5] END ..C=1, gamma=scale, kernel=sigmoid;, score=0.401 total time= 3.7s
[CV 3/5] END ....C=1, gamma=auto, kernel=linear;, score=0.816 total time= 1.8s
[CV 1/5] END ....C=1, gamma=auto, kernel=linear;, score=0.820 total time= 2.8s
[CV 2/5] END ....C=1, gamma=auto, kernel=linear;, score=0.841 total time= 2.8s
[CV 4/5] END ....C=1, gamma=auto, kernel=linear;, score=0.823 total time= 2.5s
[CV 5/5] END ....C=1, gamma=auto, kernel=linear;, score=0.828 total time= 2.2s
[CV 2/5] END ......C=1, gamma=auto, kernel=poly;, score=0.439 total time= 7.6s
[CV 4/5] END ......C=1, gamma=auto, kernel=poly;, score=0.431 total time= 7.5s
[CV 1/5] END ......C=1, gamma=auto, kernel=poly;, score=0.430 total time= 11.7s
[CV 3/5] END ......C=1, gamma=auto, kernel=poly;, score=0.432 total time= 10.6s
[CV 1/5] END .......C=1, gamma=auto, kernel=rbf;, score=0.805 total time= 5.1s
[CV 2/5] END .......C=1, gamma=auto, kernel=rbf;, score=0.792 total time= 10.3s
[CV 3/5] END .......C=1, gamma=auto, kernel=rbf;, score=0.772 total time= 10.4s
[CV 5/5] END ......C=1, gamma=auto, kernel=poly;, score=0.427 total time= 14.7s
[CV 4/5] END .......C=1, gamma=auto, kernel=rbf;, score=0.796 total time= 12.9s
[CV 5/5] END .......C=1, gamma=auto, kernel=rbf;, score=0.774 total time= 14.2s
[CV 1/5] END ...C=1, gamma=auto, kernel=sigmoid;, score=0.770 total time= 14.1s
[CV 2/5] END ...C=1, gamma=auto, kernel=sigmoid;, score=0.782 total time= 16.1s
[CV 3/5] END ...C=1, gamma=auto, kernel=sigmoid;, score=0.752 total time= 13.1s
[CV 1/5] END ...C=2, gamma=scale, kernel=linear;, score=0.812 total time= 6.5s
[CV 2/5] END ...C=2, gamma=scale, kernel=linear;, score=0.836 total time= 7.5s
[CV 4/5] END ...C=1, gamma=auto, kernel=sigmoid;, score=0.764 total time= 12.2s
[CV 5/5] END ...C=1, gamma=auto, kernel=sigmoid;, score=0.759 total time= 15.0s
[CV 3/5] END ...C=2, gamma=scale, kernel=linear;, score=0.814 total time= 7.0s
[CV 5/5] END ...C=2, gamma=scale, kernel=linear;, score=0.823 total time= 5.8s
[CV 4/5] END ...C=2, gamma=scale, kernel=linear;, score=0.829 total time= 6.9s
[CV 1/5] END .....C=2, gamma=scale, kernel=poly;, score=0.805 total time= 7.6s
[CV 2/5] END .....C=2, gamma=scale, kernel=poly;, score=0.811 total time= 8.2s
[CV 3/5] END .....C=2, gamma=scale, kernel=poly;, score=0.797 total time= 7.0s
[CV 4/5] END .....C=2, gamma=scale, kernel=poly;, score=0.799 total time= 6.6s
[CV 5/5] END .....C=2, gamma=scale, kernel=poly;, score=0.809 total time= 2.9s
[CV 3/5] END ......C=2, gamma=scale, kernel=rbf;, score=0.844 total time= 2.9s
[CV 1/5] END ......C=2, gamma=scale, kernel=rbf;, score=0.856 total time= 4.5s
[CV 4/5] END ......C=2, gamma=scale, kernel=rbf;, score=0.860 total time= 2.9s
[CV 2/5] END ......C=2, gamma=scale, kernel=rbf;, score=0.864 total time= 4.6s
[CV 5/5] END ......C=2, gamma=scale, kernel=rbf;, score=0.861 total time= 2.7s
[CV 2/5] END ..C=2, gamma=scale, kernel=sigmoid;, score=0.394 total time= 3.0s
[CV 4/5] END ..C=2, gamma=scale, kernel=sigmoid;, score=0.449 total time= 3.3s
[CV 1/5] END ..C=2, gamma=scale, kernel=sigmoid;, score=0.396 total time= 5.5s
[CV 3/5] END ..C=2, gamma=scale, kernel=sigmoid;, score=0.409 total time= 5.5s
[CV 5/5] END ..C=2, gamma=scale, kernel=sigmoid;, score=0.376 total time= 3.2s
[CV 1/5] END ....C=2, gamma=auto, kernel=linear;, score=0.812 total time= 1.6s
[CV 2/5] END ....C=2, gamma=auto, kernel=linear;, score=0.836 total time= 2.4s
[CV 4/5] END ....C=2, gamma=auto, kernel=linear;, score=0.829 total time= 1.8s
[CV 3/5] END ....C=2, gamma=auto, kernel=linear;, score=0.814 total time= 2.6s
[CV 5/5] END ....C=2, gamma=auto, kernel=linear;, score=0.823 total time= 3.0s
[CV 1/5] END ......C=2, gamma=auto, kernel=poly;, score=0.510 total time= 7.2s
[CV 2/5] END ......C=2, gamma=auto, kernel=poly;, score=0.562 total time= 7.7s
[CV 4/5] END ......C=2, gamma=auto, kernel=poly;, score=0.514 total time= 7.6s
[CV 3/5] END ......C=2, gamma=auto, kernel=poly;, score=0.544 total time= 10.0s
[CV 1/5] END .......C=2, gamma=auto, kernel=rbf;, score=0.820 total time= 3.6s
[CV 2/5] END .......C=2, gamma=auto, kernel=rbf;, score=0.809 total time= 4.0s
[CV 5/5] END ......C=2, gamma=auto, kernel=poly;, score=0.534 total time= 6.9s
[CV 3/5] END .......C=2, gamma=auto, kernel=rbf;, score=0.796 total time= 3.9s
[CV 4/5] END .......C=2, gamma=auto, kernel=rbf;, score=0.806 total time= 3.2s
[CV 5/5] END .......C=2, gamma=auto, kernel=rbf;, score=0.787 total time= 4.5s
[CV 3/5] END ...C=2, gamma=auto, kernel=sigmoid;, score=0.776 total time= 6.1s
[CV 1/5] END ...C=2, gamma=auto, kernel=sigmoid;, score=0.801 total time= 7.0s
[CV 2/5] END ...C=2, gamma=auto, kernel=sigmoid;, score=0.792 total time= 8.2s
[CV 4/5] END ...C=2, gamma=auto, kernel=sigmoid;, score=0.794 total time= 9.3s
[CV 1/5] END ...C=3, gamma=scale, kernel=linear;, score=0.814 total time= 6.5s
[CV 2/5] END ...C=3, gamma=scale, kernel=linear;, score=0.838 total time= 5.2s
[CV 5/5] END ...C=2, gamma=auto, kernel=sigmoid;, score=0.775 total time= 9.4s
[CV 3/5] END ...C=3, gamma=scale, kernel=linear;, score=0.810 total time= 4.8s
[CV 4/5] END ...C=3, gamma=scale, kernel=linear;, score=0.828 total time= 5.8s
[CV 5/5] END ...C=3, gamma=scale, kernel=linear;, score=0.820 total time= 6.2s
[CV 1/5] END .....C=3, gamma=scale, kernel=poly;, score=0.815 total time= 4.8s
[CV 2/5] END .....C=3, gamma=scale, kernel=poly;, score=0.820 total time= 6.8s
[CV 3/5] END .....C=3, gamma=scale, kernel=poly;, score=0.806 total time= 5.6s
[CV 4/5] END .....C=3, gamma=scale, kernel=poly;, score=0.805 total time= 5.5s
[CV 5/5] END .....C=3, gamma=scale, kernel=poly;, score=0.816 total time= 5.8s
[CV 1/5] END ......C=3, gamma=scale, kernel=rbf;, score=0.859 total time= 8.2s
[CV 2/5] END ......C=3, gamma=scale, kernel=rbf;, score=0.864 total time= 8.6s
[CV 3/5] END ......C=3, gamma=scale, kernel=rbf;, score=0.845 total time= 8.6s
[CV 4/5] END ......C=3, gamma=scale, kernel=rbf;, score=0.860 total time= 7.8s
[CV 1/5] END ..C=3, gamma=scale, kernel=sigmoid;, score=0.386 total time= 8.5s
[CV 3/5] END ..C=3, gamma=scale, kernel=sigmoid;, score=0.401 total time= 7.6s
[CV 5/5] END ......C=3, gamma=scale, kernel=rbf;, score=0.869 total time= 9.8s
[CV 2/5] END ..C=3, gamma=scale, kernel=sigmoid;, score=0.388 total time= 10.0s
[CV 1/5] END ....C=3, gamma=auto, kernel=linear;, score=0.814 total time= 5.7s
[CV 2/5] END ....C=3, gamma=auto, kernel=linear;, score=0.838 total time= 6.3s
[CV 5/5] END ..C=3, gamma=scale, kernel=sigmoid;, score=0.379 total time= 9.2s
[CV 4/5] END ..C=3, gamma=scale, kernel=sigmoid;, score=0.450 total time= 9.3s
[CV 3/5] END ....C=3, gamma=auto, kernel=linear;, score=0.810 total time= 5.5s
[CV 4/5] END ....C=3, gamma=auto, kernel=linear;, score=0.828 total time= 6.7s
[CV 5/5] END ....C=3, gamma=auto, kernel=linear;, score=0.820 total time= 6.2s
[CV 1/5] END ......C=3, gamma=auto, kernel=poly;, score=0.562 total time= 11.1s
[CV 2/5] END ......C=3, gamma=auto, kernel=poly;, score=0.546 total time= 9.6s
[CV 3/5] END ......C=3, gamma=auto, kernel=poly;, score=0.557 total time= 8.5s
[CV 4/5] END ......C=3, gamma=auto, kernel=poly;, score=0.532 total time= 8.4s
[CV 1/5] END .......C=3, gamma=auto, kernel=rbf;, score=0.831 total time= 5.4s
[CV 3/5] END .......C=3, gamma=auto, kernel=rbf;, score=0.804 total time= 3.5s
[CV 5/5] END ......C=3, gamma=auto, kernel=poly;, score=0.542 total time= 7.0s
[CV 2/5] END .......C=3, gamma=auto, kernel=rbf;, score=0.825 total time= 4.3s
[CV 5/5] END .......C=3, gamma=auto, kernel=rbf;, score=0.810 total time= 3.0s
[CV 1/5] END ...C=3, gamma=auto, kernel=sigmoid;, score=0.810 total time= 3.0s
[CV 4/5] END .......C=3, gamma=auto, kernel=rbf;, score=0.823 total time= 4.8s
[CV 2/5] END ...C=3, gamma=auto, kernel=sigmoid;, score=0.801 total time= 4.3s
[CV 3/5] END ...C=3, gamma=auto, kernel=sigmoid;, score=0.784 total time= 3.2s
[CV 4/5] END ...C=3, gamma=auto, kernel=sigmoid;, score=0.805 total time= 3.2s
[CV 1/5] END ...C=4, gamma=scale, kernel=linear;, score=0.814 total time= 2.9s
[CV 2/5] END ...C=4, gamma=scale, kernel=linear;, score=0.833 total time= 1.9s
[CV 3/5] END ...C=4, gamma=scale, kernel=linear;, score=0.810 total time= 1.9s
[CV 5/5] END ...C=3, gamma=auto, kernel=sigmoid;, score=0.784 total time= 4.2s
[CV 1/5] END .....C=4, gamma=scale, kernel=poly;, score=0.820 total time= 2.0s
[CV 4/5] END ...C=4, gamma=scale, kernel=linear;, score=0.821 total time= 2.4s
[CV 5/5] END ...C=4, gamma=scale, kernel=linear;, score=0.819 total time= 2.1s
[CV 2/5] END .....C=4, gamma=scale, kernel=poly;, score=0.830 total time= 3.5s
[CV 3/5] END .....C=4, gamma=scale, kernel=poly;, score=0.812 total time= 2.1s
[CV 5/5] END .....C=4, gamma=scale, kernel=poly;, score=0.828 total time= 2.1s
[CV 4/5] END .....C=4, gamma=scale, kernel=poly;, score=0.801 total time= 3.1s
[CV 1/5] END ......C=4, gamma=scale, kernel=rbf;, score=0.861 total time= 3.4s
[CV 3/5] END ......C=4, gamma=scale, kernel=rbf;, score=0.849 total time= 3.3s
[CV 2/5] END ......C=4, gamma=scale, kernel=rbf;, score=0.868 total time= 3.5s
[CV 4/5] END ......C=4, gamma=scale, kernel=rbf;, score=0.864 total time= 3.3s
[CV 1/5] END ..C=4, gamma=scale, kernel=sigmoid;, score=0.384 total time= 3.5s
[CV 2/5] END ..C=4, gamma=scale, kernel=sigmoid;, score=0.400 total time= 4.0s
[CV 5/5] END ......C=4, gamma=scale, kernel=rbf;, score=0.870 total time= 4.2s
[CV 3/5] END ..C=4, gamma=scale, kernel=sigmoid;, score=0.396 total time= 4.2s
[CV 1/5] END ....C=4, gamma=auto, kernel=linear;, score=0.814 total time= 5.8s
[CV 2/5] END ....C=4, gamma=auto, kernel=linear;, score=0.833 total time= 6.9s
[CV 4/5] END ..C=4, gamma=scale, kernel=sigmoid;, score=0.444 total time= 8.8s
[CV 5/5] END ..C=4, gamma=scale, kernel=sigmoid;, score=0.376 total time= 10.3s
[CV 3/5] END ....C=4, gamma=auto, kernel=linear;, score=0.810 total time= 6.0s
[CV 5/5] END ....C=4, gamma=auto, kernel=linear;, score=0.819 total time= 5.2s
[CV 4/5] END ....C=4, gamma=auto, kernel=linear;, score=0.821 total time= 6.0s
[CV 1/5] END ......C=4, gamma=auto, kernel=poly;, score=0.574 total time= 16.4s
[CV 2/5] END ......C=4, gamma=auto, kernel=poly;, score=0.568 total time= 16.7s
[CV 3/5] END ......C=4, gamma=auto, kernel=poly;, score=0.580 total time= 15.8s
[CV 4/5] END ......C=4, gamma=auto, kernel=poly;, score=0.554 total time= 19.0s
[CV 2/5] END .......C=4, gamma=auto, kernel=rbf;, score=0.830 total time= 7.4s
[CV 1/5] END .......C=4, gamma=auto, kernel=rbf;, score=0.831 total time= 9.8s
[CV 3/5] END .......C=4, gamma=auto, kernel=rbf;, score=0.807 total time= 9.3s
[CV 5/5] END ......C=4, gamma=auto, kernel=poly;, score=0.576 total time= 16.8s
[CV 4/5] END .......C=4, gamma=auto, kernel=rbf;, score=0.826 total time= 8.4s
[CV 5/5] END .......C=4, gamma=auto, kernel=rbf;, score=0.821 total time= 8.9s
[CV 1/5] END ...C=4, gamma=auto, kernel=sigmoid;, score=0.818 total time= 7.2s
[CV 2/5] END ...C=4, gamma=auto, kernel=sigmoid;, score=0.809 total time= 8.5s
[CV 3/5] END ...C=4, gamma=auto, kernel=sigmoid;, score=0.789 total time= 7.2s
[CV 4/5] END ...C=4, gamma=auto, kernel=sigmoid;, score=0.810 total time= 6.7s
[CV 5/5] END ...C=4, gamma=auto, kernel=sigmoid;, score=0.790 total time= 5.0s
# Best Model
# Extract the attributes of the best model.
#===========================Fill This===========================#
# 1. Extract the best score.
# 2. Extract a dictionary of the parameters.
# !! Use the attributes of the `oGsSvc` object.
bestScore = oGsSvc.best_score_
dBestParams = oGsSvc.best_params_
#===============================================================#
print(f'The best model had the following parameters: {dBestParams} with the CV score: {bestScore:0.2%}')
The best model had the following parameters: {'C': 4, 'gamma': 'scale', 'kernel': 'rbf'} with the CV score: 86.23%
(#) In production one would visualize the effect of each parameter on the model result. Then use it to fine tune farther the parameters.
oGsSvc.score(mXTest, vYTest) ## will be the same like extracting best model
0.85
# The Best Model
#===========================Fill This===========================#
# 1. Extract the best model.
# 2. Score the best model on the test data set.
bestModel = oGsSvc.best_estimator_
modelScore = bestModel.score(mXTest, vYTest)
#===============================================================#
print(f'The model score (Accuracy) on the data: {modelScore:0.2%}') #<! Accuracy
The model score (Accuracy) on the data: 85.00%
(#) With proper tuning one can improve the baseline model by
~5%.
Train the Best Model on the Train Data Set#
In production we take the optimal Hyper Parameters and then retrain the model on the whole training data set.
This is the model we’ll use in production.
# The Model with Optimal Parameters
#===========================Fill This===========================#
# 1. Construct the model.
# 2. Train the model.
oSvmCls = bestModel.fit(mXTrain, vYTrain)
#===============================================================#
modelScore = oSvmCls.score(mXTest, vYTest)
print(f'The model score (Accuracy) on the data: {modelScore:0.2%}') #<! Accuracy
The model score (Accuracy) on the data: 85.00%
(?) Is the value above exactly as the value from the best model of the grid search? If so, look at the
refitparameter ofGridSearchCV.
Performance Metrics / Scores#
In this section we’ll analyze the model using the confusion matrix.
Plot the Confusion Matrix#
# Plot the Confusion Matrix
hF, hA = plt.subplots(figsize = (10, 10))
#===========================Fill This===========================#
# 1. Plot the confusion matrix for the best model.
# 2. Use the data labels (`L_CLASSES`).
hA, mConfMat = PlotConfusionMatrix(vYTest, oSvmCls.predict(mXTest),lLabels= L_CLASSES, hA = hA)
#===============================================================#
plt.show()
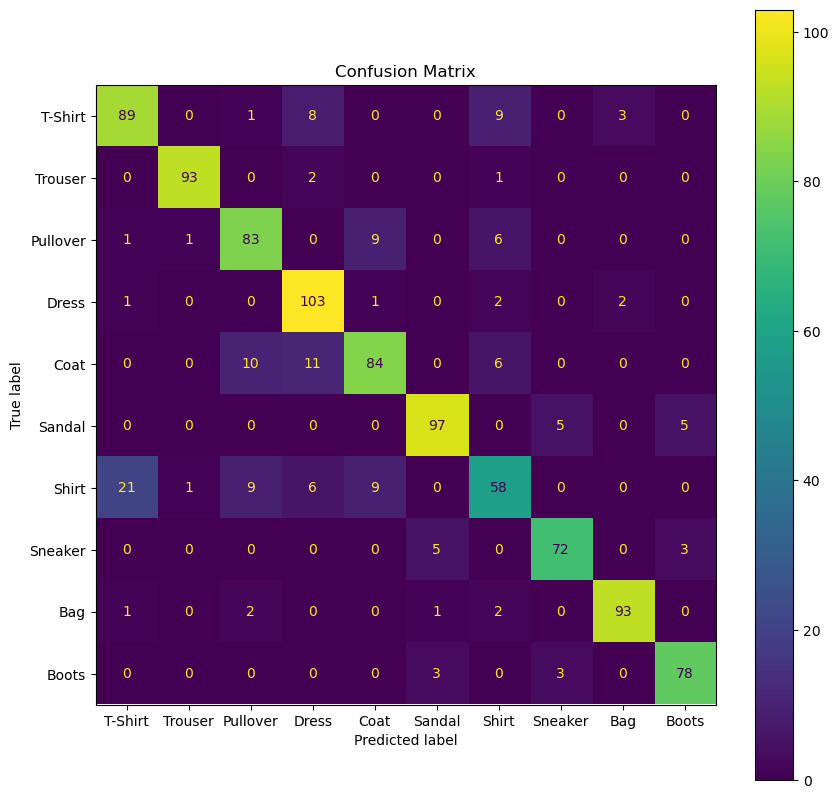
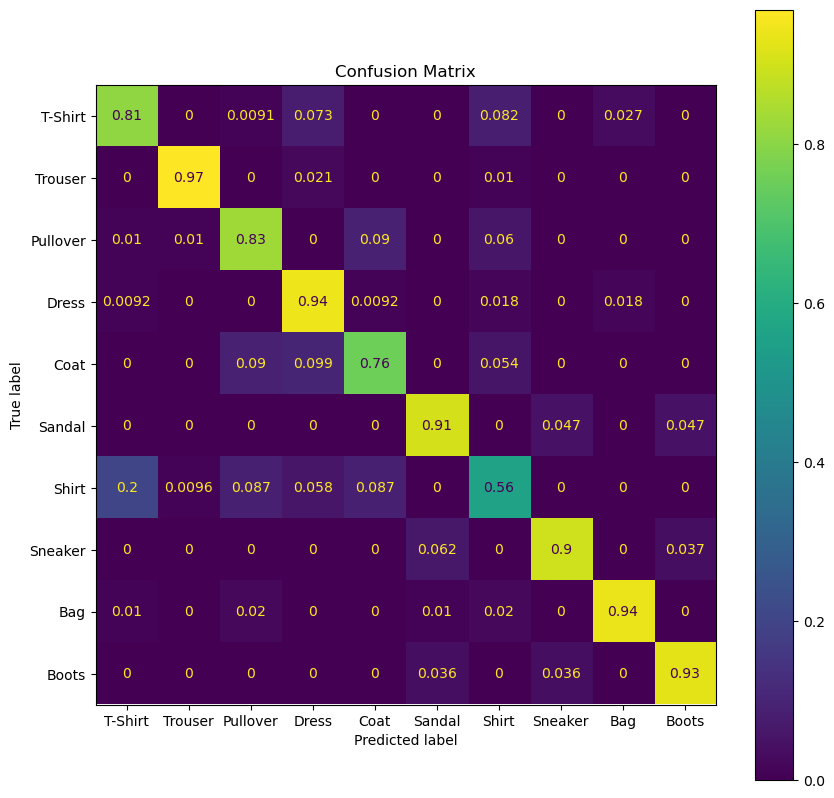
(?) Which class has the best accuracy?
(?) Which class has a dominant false prediction? Does it make sense?
(?) What’s the difference between \(p \left( \hat{y}_{i} = \text{coat} \mid {x}_{i} = \text{coat} \right)\) to \(p \left( {y}_{i} = \text{coat} \mid \hat{y}_{i} = \text{coat} \right)\)?
recall and presicion
(!) Make the proper calculations on
mConfMator the functionPlotConfusionMatrixto answer the questions above.
# Plot the Confusion Matrix
hF, hA = plt.subplots(figsize = (10, 10))
#===========================Fill This===========================#
# 1. Plot the confusion matrix for the best model.
# 2. Use the data labels (`L_CLASSES`).
hA, mConfMat = PlotConfusionMatrix(vYTest, oSvmCls.predict(mXTest),lLabels= L_CLASSES,normMethod= 'true' , hA = hA)
#===============================================================#
plt.show()