MultiDimensional Scaling (MDS)#
Notebook by:
Royi Avital RoyiAvital@fixelalgorithms.com
Revision History#
Version |
Date |
User |
Content / Changes |
|---|---|---|---|
1.0.000 |
13/04/2024 |
Royi Avital |
First version |
# Import Packages
# General Tools
import numpy as np
import scipy as sp
import pandas as pd
# Machine Learning
from sklearn.datasets import make_s_curve
from sklearn.manifold import MDS
# Miscellaneous
import math
import os
from platform import python_version
import random
import timeit
# Typing
from typing import Callable, Dict, List, Optional, Self, Set, Tuple, Union
# Visualization
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
# Jupyter
from IPython import get_ipython
from IPython.display import Image
from IPython.display import display
from ipywidgets import Dropdown, FloatSlider, interact, IntSlider, Layout, SelectionSlider
from ipywidgets import interact
Notations#
(?) Question to answer interactively.
(!) Simple task to add code for the notebook.
(@) Optional / Extra self practice.
(#) Note / Useful resource / Food for thought.
Code Notations:
someVar = 2; #<! Notation for a variable
vVector = np.random.rand(4) #<! Notation for 1D array
mMatrix = np.random.rand(4, 3) #<! Notation for 2D array
tTensor = np.random.rand(4, 3, 2, 3) #<! Notation for nD array (Tensor)
tuTuple = (1, 2, 3) #<! Notation for a tuple
lList = [1, 2, 3] #<! Notation for a list
dDict = {1: 3, 2: 2, 3: 1} #<! Notation for a dictionary
oObj = MyClass() #<! Notation for an object
dfData = pd.DataFrame() #<! Notation for a data frame
dsData = pd.Series() #<! Notation for a series
hObj = plt.Axes() #<! Notation for an object / handler / function handler
Code Exercise#
Single line fill
vallToFill = ???
Multi Line to Fill (At least one)
# You need to start writing
????
Section to Fill
#===========================Fill This===========================#
# 1. Explanation about what to do.
# !! Remarks to follow / take under consideration.
mX = ???
???
#===============================================================#
# Configuration
# %matplotlib inline
seedNum = 512
np.random.seed(seedNum)
random.seed(seedNum)
# Matplotlib default color palette
lMatPltLibclr = ['#1f77b4', '#ff7f0e', '#2ca02c', '#d62728', '#9467bd', '#8c564b', '#e377c2', '#7f7f7f', '#bcbd22', '#17becf']
# sns.set_theme() #>! Apply SeaBorn theme
runInGoogleColab = 'google.colab' in str(get_ipython())
# Constants
FIG_SIZE_DEF = (8, 8)
ELM_SIZE_DEF = 50
CLASS_COLOR = ('b', 'r')
EDGE_COLOR = 'k'
MARKER_SIZE_DEF = 10
LINE_WIDTH_DEF = 2
# Courses Packages
import sys
sys.path.append('../')
sys.path.append('../../')
sys.path.append('../../../')
from utils.DataVisualization import PlotScatterData, PlotScatterData3D
# General Auxiliary Functions
Dimensionality Reduction by MDS#
The MDS is a non linear transformation from \(\mathbb{R}^{D} \to \mathbb{R}^{d}\) where \(d \ll D\).
Given a set \(\mathcal{X} = {\left\{ \boldsymbol{x}_{i} \in \mathbb{R}^{D} \right\}}_{i = 1}^{n}\) is builds the set \(\mathcal{Z} = {\left\{ \boldsymbol{z}_{i} \in \mathbb{R}^{d} \right\}}_{i = 1}^{n}\) such that the distance matrices of each set are similar.
In this notebook:
We’ll implement the classic MDS.
We’ll use the data set to show the effects of dimensionality reduction.
# Parameters
# Data
numSamples = 1000
# Model
lowDim = 2
Generate / Load Data#
In this notebook we’ll use SciKit Learn’s make_s_curve to generated data.
# Generate Data
mX, vC = make_s_curve(numSamples) #<! Results are random beyond the noise
print(f'The features data shape: {mX.shape}')
print(f'The features data type: {mX.dtype}')
The features data shape: (1000, 3)
The features data type: float64
Plot Data#
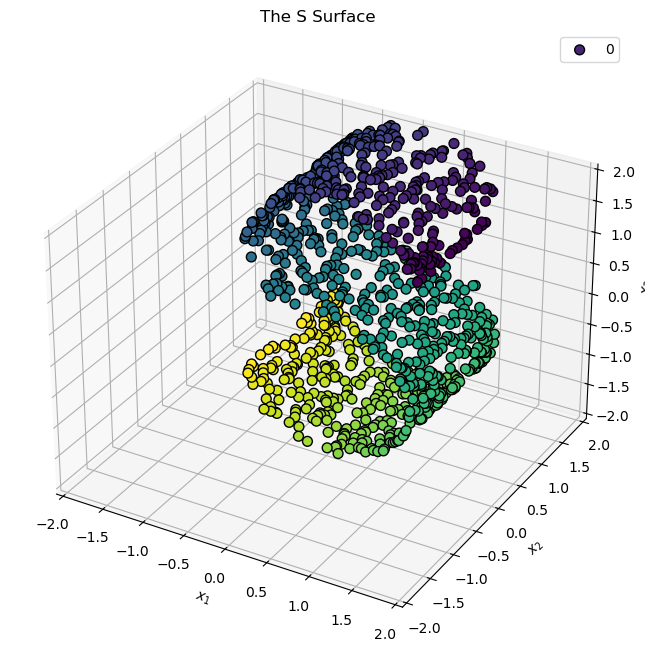
# Plot the Data
hA = PlotScatterData3D(mX, vC = vC)
hA.set_xlim([-2, 2])
hA.set_ylim([-2, 2])
hA.set_zlim([-2, 2])
hA.set_title('The S Surface')
plt.show()
Applying Dimensionality Reduction - MDS#
In this section we’ll implement the Classic MDs:
Non Metric (Classic) MDS#
set \(\boldsymbol{K}_{x}=-\frac{1}{2}\boldsymbol{J}\boldsymbol{D}_{x}\boldsymbol{J}\)
where \(\boldsymbol{J}=\left(\boldsymbol{I}-\frac{1}{N}\boldsymbol{1}\boldsymbol{1}^{T}\right)\)Decompose \(\boldsymbol{K}_{x}=\boldsymbol{W}\boldsymbol{\Lambda}\boldsymbol{W}\)
set \(\boldsymbol{Z}=\boldsymbol{\Lambda}_{d}^{\frac{1}{2}}\boldsymbol{W}_{d}^{T}\)
(#) The non metric MDS matches (Kernel) PCA.
(#) It is assumed above that the eigen values matrix \(\boldsymbol{\Lambda}\) is sorted.
(#) For Euclidean distance there is a closed form solution (As with the PCA).
# Classic MDS Implementation
def ClassicalMDS( mD: np.ndarray, lowDim: int ) -> np.ndarray:
numSamples = mD.shape[0]
mJ = np.eye(numSamples) - ((1 / numSamples) * np.ones((numSamples, numSamples)))
mK = -0.5 * mJ @ mD @ mJ #<! Due to the form of mJ one can avoid the matrix multiplication
vL, mW = np.linalg.eigh(mK)
# Sort Eigen Values
vIdx = np.argsort(-vL)
vL = vL[vIdx]
mW = mW[:, vIdx]
# Reconstruct
mZ = mW[:, :lowDim] * np.sqrt(vL[:lowDim])
return mZ
# Apply the MDS
# Build the Distance Matrix
mD = sp.spatial.distance.squareform(sp.spatial.distance.pdist(mX))
# The MDS output
mZ1 = ClassicalMDS(np.square(mD), lowDim)
(?) Could we achieve the above using a different method?
# Plot the Low Dimensional Data
hA = PlotScatterData3D(mZ1, vC = vC, axesProjection = None, figSize = (8, 8))
hA.set_xlabel('${{z}}_{{1}}$')
hA.set_ylabel('${{z}}_{{2}}$')
hA.set_box_aspect(1)
hA.set_title('Classical Euclidean MDS = PCA')
plt.show()
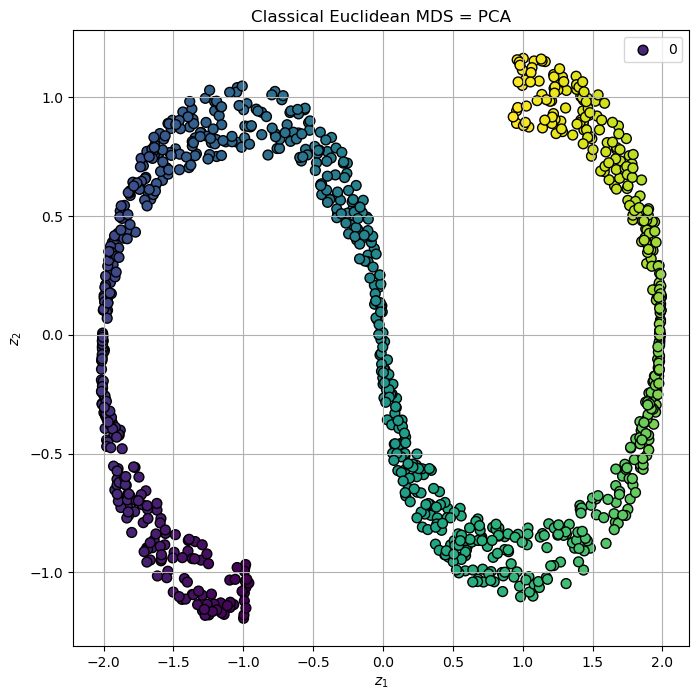
(#) The Non Metric MDS is guaranteed to keep the order of distances, but not the distance itself.
Metric MDS#
the above is not deterministic (It is a non convex optimization problem)!
# Apply MDS using SciKit Learn
oMdsDr = MDS(n_components = lowDim, dissimilarity = 'precomputed', normalized_stress = 'auto')
mZ2 = oMdsDr.fit_transform(mD)
(?) Are the results deterministic in the case above?
(#) The Metric MDS tries to rebuild the data in low dimension with as similar as it can distance matrix. Yet it is not guaranteed to have the same distance.
# Plot the Low Dimensional Data
hA = PlotScatterData3D(mZ2, vC = vC, axesProjection = None, figSize = (8, 8))
hA.set_xlabel('${{z}}_{{1}}$')
hA.set_ylabel('${{z}}_{{2}}$')
hA.set_box_aspect(1)
hA.set_title(r'Metric Euclidean MDS $\neq$ PCA')
plt.show()
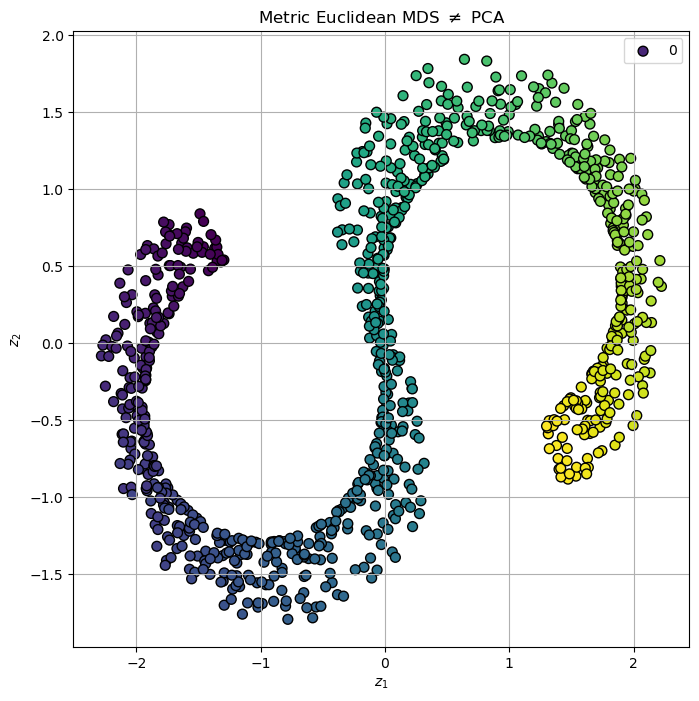
Metric MDS with Geodesic Distance#
We have access to the geodesic distance using vC, the position along the “main” axis.
# Geodesic Distance
mGeodesicDist = sp.spatial.distance.squareform(sp.spatial.distance.pdist(np.c_[vC, mX[:, 1]]))
mZ3 = oMdsDr.fit_transform(mGeodesicDist)
(?) Can we use the geodesic distance in real world?
# Plot the Low Dimensional Data
hA = PlotScatterData3D(mZ3, vC = vC, axesProjection = None, figSize = (8, 8))
hA.set_xlabel('${{z}}_{{1}}$')
hA.set_ylabel('${{z}}_{{2}}$')
hA.set_box_aspect(1)
hA.set_title(r'Metric Geodesic MDS $\neq$ PCA')
plt.show()
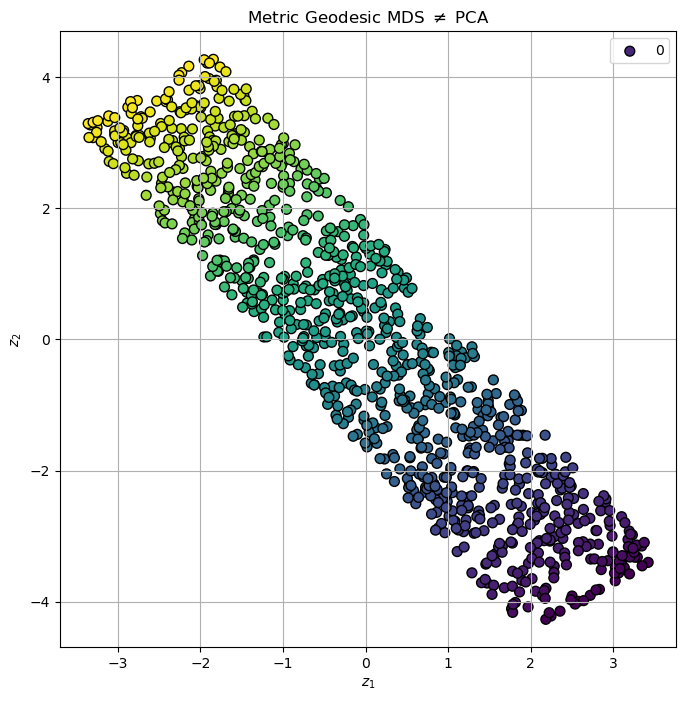
(?) Is the result above better than the previous ones? Why?
